Multi-Omics and Single-Cell Mendelian Randomization Reveal a Potential Role of VNN2 in Lung Adenocarcinoma in Resting Natural Killer Cells
DOI:
https://doi.org/10.14740/wjon2689Keywords:
VNN2, Lung adenocarcinoma, Mendelian randomization, sc-eQTL, Resting NK cellsAbstract
Background: We aimed to evaluate the potential association between genetically predicted vanin-2 (VNN2) expression and lung adenocarcinoma (LUAD) risk, and to explore the immune cell subtype that may underlie this relationship.
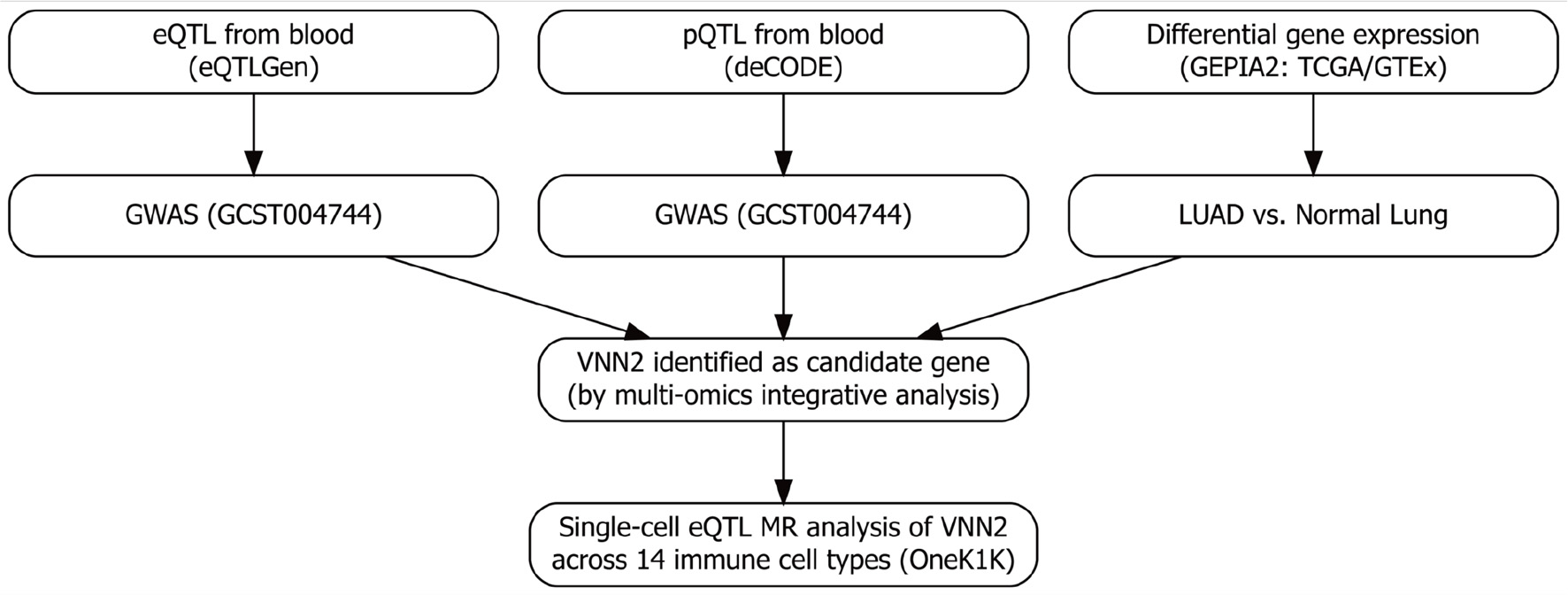
Methods: We integrated whole-blood expression quantitative trait loci (eQTL) data from eQTLGen, plasma protein quantitative trait loci (pQTL) data from deCODE, and LUAD genome-wide association study (GWAS) data from European-ancestry cohorts, together with differential expression analysis using GEPIA2, to identify candidate genes for subsequent single-cell eQTL (sc-eQTL) Mendelian randomization (MR) analysis. For the sc-eQTL analysis, VNN2-associated eQTLs from 14 immune cell types profiled in the OneK1K single-cell eQTL resource were tested for associations with LUAD risk.
Results: Bulk-level MR analysis showed that genetically predicted increases in VNN2 expression and protein levels were significantly associated with a reduced risk of LUAD (eQTL-MR: odds ratio (OR) = 0.964, 95% confidence interval (95% CI), 0.934–0.995; P = 0.024; pQTL-MR: OR = 0.946, 95% CI, 0.921–0.970; P = 2.87 × 10−5). Transcriptomic analyses confirmed significant downregulation of VNN2 in LUAD tumors compared with normal lung tissues. sc-eQTL MR identified the strongest association in resting natural killer (rNK) cells (OR = 0.896, 95% CI, 0.829–0.967; P = 0.005).
Conclusions: Multi-omics and sc-eQTL MR analyses indicated that genetically predicted increases in VNN2 expression were associated with a reduced risk of LUAD, with the most pronounced effect observed in rNK cells. These findings suggest a potential cell type–specific role of VNN2 in LUAD susceptibility and warrant further studies to validate its biological relevance and clinical implications.
Published
Issue
Section
License
Copyright (c) 2026 The authors

This work is licensed under a Creative Commons Attribution 4.0 International License.









