Pan-Cancer Analysis of NOP2 Reveals Its Prognostic Relevance and Association With the Tumor Immune Microenvironment
DOI:
https://doi.org/10.14740/wjon2739Keywords:
NOP2, Pan-cancer, Prognosis, Immune infiltration, BioinformaticsAbstract
Background: RNA 5-methylcytosine (m5C) modification has emerged as an important layer of epigenetic regulation in cancer biology. NOP2 (also known as NSUN1), a nucleolar RNA methyltransferase involved in ribosomal RNA methylation and ribosome biogenesis, has been reported to promote tumor progression in several individual malignancies. However, the expression pattern, prognostic relevance, and immune-related features across multiple cancer types remain incompletely understood.
Methods: We conducted an integrative pan-cancer analysis of NOP2 across 33 tumor types using publicly available datasets and web tools, including The Cancer Genome Atlas (TCGA), the Human Protein Atlas (HPA), Gene Expression Profiling Interactive Analysis 2 (GEPIA2), cBioPortal for Cancer Genomics (cBioPortal), the University of Alabama at Birmingham Cancer data analysis portal (UALCAN), Search Tool for the Retrieval of Interacting Genes/Proteins (STRING), and SangerBox. NOP2 expression differences between tumor and normal tissues were evaluated at both mRNA and protein levels. Survival analyses were conducted using GEPIA2, and genetic alterations were characterized via cBioPortal. The relationships between NOP2 expression and the tumor immune microenvironment were assessed by estimating tumor-infiltrating immune cell proportions with Cell-type Identification By Estimating Relative Subsets Of RNA Transcripts (CIBERSORT), followed by correlation analyses with immune cell infiltration levels, immune checkpoint-related genes, tumor mutational burden (TMB), and microsatellite instability (MSI). Finally, protein-protein interaction and co-expression analyses were conducted to identify NOP2-interacting and NOP2-correlated genes, which were subjected to functional enrichment analyses to infer potential biological pathways associated with NOP2.
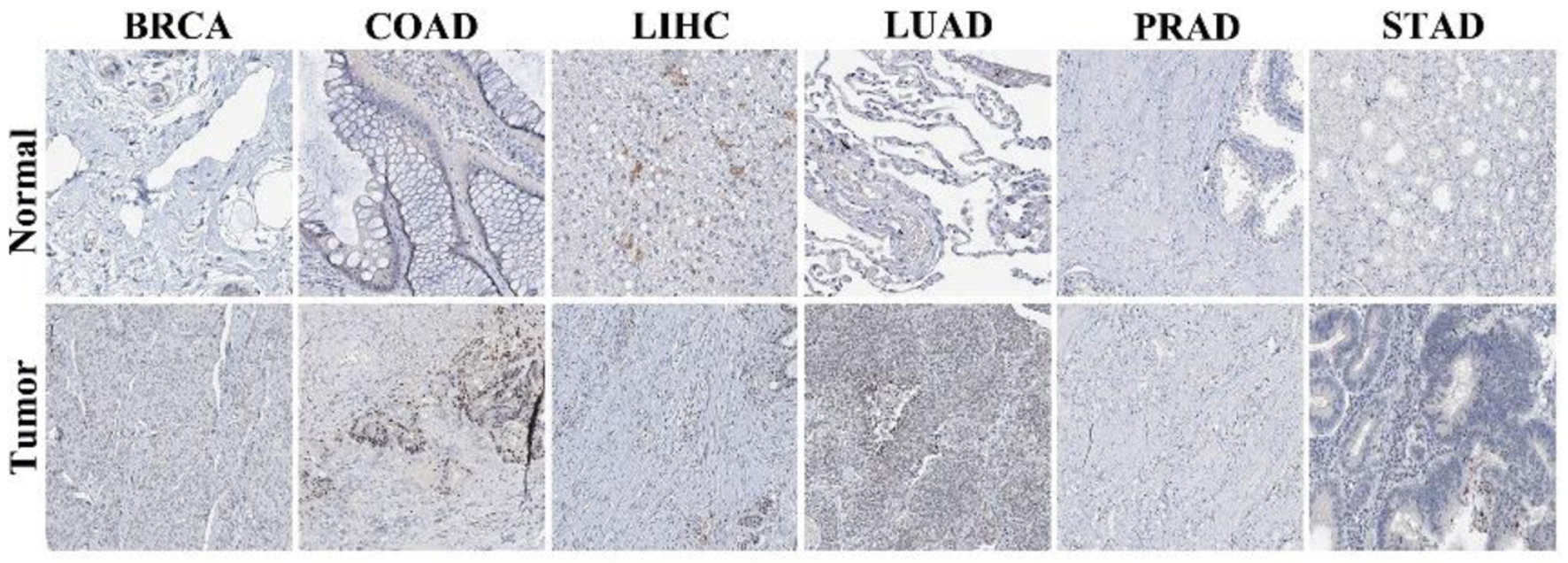
Results: NOP2 mRNA was significantly upregulated in 20 of 33 tumor types compared with normal tissues (|log2FC| ≥ 1, q < 0.01), with protein-level overexpression confirmed in most cancers examined (P < 0.001). High NOP2 expression was associated with shorter overall survival (OS) in 11 cancer types and poorer disease-free survival (DFS) in 13 cancer types (P < 0.05), including renal cell carcinomas, hepatocellular carcinoma, lower-grade glioma, and lung adenocarcinoma. Genetic alterations of NOP2 were most frequent in ovarian cancer (6.1%) and patients with NOP2 genetic alterations were associated with poorer OS, DSS, and PFS in the cBioPortal cohort analysis. NOP2 expression correlated significantly with mast cell infiltration in 24 cancer types and CD8+ T-cell infiltration in 19 cancer types (P < 0.05). Positive correlations between NOP2 expression and TMB were observed in 14 of 15 cancer types showing significant associations. Functional enrichment analysis confirmed associations with ribosome biogenesis and RNA processing pathways.
Conclusion: This pan-cancer analysis shows that NOP2 is broadly dysregulated and associated with adverse survival and immune-related features across multiple human cancers. However, given the exploratory and largely univariate nature of the present study, its independent prognostic value and clinical utility require further validation in cancer-specific cohorts with multivariable and comparative predictive analyses.
Published
Issue
Section
License
Copyright (c) 2026 The authors

This work is licensed under a Creative Commons Attribution 4.0 International License.









