| World Journal of Oncology, ISSN 1920-4531 print, 1920-454X online, Open Access |
| Article copyright, the authors; Journal compilation copyright, World J Oncol and Elmer Press Inc |
| Journal website https://wjon.elmerpub.com |
Original Article
Volume 16, Number 2, April 2025, pages 182-199
Increased Sirtuin 6 Activity in Tumor Cells Can Prompt CD4-Positive T-Cell Differentiation Into Regulatory T Cells and Impede Immune Surveillance in the Microenvironment
Nan Yang Zhanga, Wen Yuan Liua, Ke Hua Fangb, c, Xiao Tian Changa, c, d
aMedical Research Center, The Affiliated Hospital of Qingdao University, Qingdao,
Shandong 266000, China
bClinical Laboratory, The Affiliated Hospital of Qingdao
University, Qingdao, Shandong 266000, China
cThese authors contributed equally
to this study.
dCorresponding Author: Xiao Tian Chang, Medical Research Center,
The Affiliated Hospital of Qingdao University, Qingdao, Shandong 266000, China
Manuscript submitted January 22, 2025, accepted March 20, 2025, published online March 25,
2025
Short title: Sirt6 in Tumor Cells Affects Immune Surveillance
doi:
https://doi.org/10.14740/wjon2547
| Abstract | ▴Top |
Background: Sirtuin 6 (Sirt6) is expressed at increased levels in many tumors and may be involved in immunoregulation. The present study investigated how Sirt6 in tumor cells affects immune surveillance.
Methods: The human tumor cell lines A2780, HeLa, Huh7, MBA-MD-231, SMMC-7721 and SW480 were incubated with UBCS039, a target-selective activator of Sirt6, to stimulate Sirt6 activity. These cells, following washing to remove residual UBCS039, were cultured with human naive CD4+ T cells in the Transwell to observe the T cell differentiation. Regulatory T cells (Tregs) among CD4+ T cells and the levels of various cytokines and adenosine (ADO), an immunosuppressive metabolite, in the culture medium, were measured via flow cytometry. The treated tumor cells were examined via transcriptomic analysis. The transcriptomic results, as well as programmed cell death protein-1 (PD-1), programmed cell death-ligand 1 (PD-L1) and Sirt6 expression in tumor cells and CD4+ T cells were verified via real-time polymerase chain reaction (PCR).
Results: Following culture with UBSC039-pretreated tumor cells, the proportion of Tregs among CD4+ T cells was significantly increased. PD-L1 and Sirt6 expressions in UBS039-pretreated tumor cells and PD-1 expression in cocultured CD4+ T cells were also increased. Moreover, the ADO level increased, and the interleukin (IL)-10, interferon (IFN)-α2, IFN-γ and monocyte chemoattractant protein-1 (MCP-1) levels decreased in the coculture medium. Transcriptomic analysis revealed significant downregulation of the antitumor genes BASP1, CPS1, GNG11, MFAP5, NNMT and SMOC1, upregulation of the tumor-promoting genes FOXA2, GSTP1, RASEF and ZNF844, and activation of adherens junctions, tumor necrosis factor (TNF)-signaling and the circadian rhythm pathway in UBCS039-pretreated SMMC-7721 cells. The above results were verified in all six cell lines.
Conclusions: The present study suggested that increased Sirt6 expression and activity in tumor cells can suppress immune surveillance by increasing Treg, ADO, PD-1 and PD-L1 levels, decreasing IFN-γ production, and altering tumor-promoting and antitumor gene expression in the microenvironment.
Keywords: Tregs; Immune surveillance; Tumor microenvironment; Sirt6; ADO; PD-1/PD-L1; IFN-γ
| Introduction | ▴Top |
Sirtuins (Sirts) are class III histone deacetylases (HDACs) that include seven members and are widely expressed in mammals [1]. Sirt6, a member of the sirtuin family, is expressed at increased levels in various tumor tissues, including hepatocellular carcinoma, ovarian cancer, renal cell carcinoma, colon carcinoma, breast cancer, osteosarcoma and prostate cancer [2-7]. Reduced Sirt6 expression was found to be related to improved overall survival [5-8]. Sirt6 is involved in multiple cancer-related signaling pathways, especially the Notch, ERK1/2 and PTEN/AKT pathways [2, 5, 9, 10]. Sirt6 also enhances epithelial-mesenchymal transition (EMT), glycolysis and oxidative phosphorylation [2, 4]. Moreover, Sirt6 silencing slowed the growth of MDA-MB-231 human breast cancer cell xenografts in transgenic mice [11]. Many studies have suggested the possible role of Sirt6 in immunoregulation, immunosenescence, immunometabolism and tumor immunity [11-15]. These studies suggest that increased Sirt6 activity and expression may facilitate tumor growth by mediating immune surveillance. However, the function of Sirt6 in tumorigenesis through regulation of the tumor immune microenvironment is poorly understood.
Regulatory T cells (Tregs) impede immune surveillance of tumors and suppress antitumor immune responses [16, 17], but the effects of tumor cells on Tregs in the tumor microenvironment (TME) are poorly understood. Tregs mainly develop from CD4+ T cells [18]. To determine how Sirt6 in tumor cells affects CD4+ T-cell differentiation in the TME, we cocultured tumor cells and human naive CD4+ T cells in the upper and lower chambers of Transwell, respectively, to mimic the TME. UBCS039 is a target-selective activator of Sirt6 that enhances the deacetylation of Sirt6-targeted histone H3 sites in human cancer cells [19, 20]. The present study used UBCS039 to treat a series of tumor cell lines, including A2780, HeLa, Huh7, MBA-MD-231, SMMC-7721 and SW480 cells, which originated from ovarian cancer, cervical cancer, liver cancer, breast tumor, liver cancer and colon cancer, respectively. The treated tumor cells were washed to remove residual UBCS039 after treatment and then cocultured with CD4+ T cells. In this study, the proportions of Tregs and programmed cell death protein-1 (PD-1) expression in cocultured CD4+ T cells, programmed cell death-ligand 1 (PD-L1) expression in pretreated tumor cells, adenosine (ADO) levels and proinflammatory cytokine production in coculture medium were measured. Furthermore, transcriptome analysis was applied to investigate the regulatory pathways of Sirt6 in UBCS039-pretreated SMMC-7721 cells. The transcriptomic results were verified in A2780, HeLa, Huh7, MBA-MD-231 and SW480 cells via real-time polymerase chain reaction (PCR). The aim of this study was to determine whether Sirt6 in tumor cells affects the tumor immune microenvironment. Our results confirmed that Sirt6 is a key gene that impedes immune surveillance to favor tumor growth.
| Materials and Methods | ▴Top |
Cell lines and culture
SMMC-7721, MBA-MD-231, MCF-7, SW480, Huh-7, HeLa and A2780 cells from various human tumors were maintained in RPMI 1640 medium (HyClone, USA) supplemented with 10% (v/v) fetal bovine serum (FBS) (Gibco, USA) and 1% (v/v) penicillin/streptomycin (Gibco) at 37 °C, with 5% CO2 in a humidified atmosphere. UBCS039 (MCE, China) was dissolved in dimethyl sulfoxide (DMSO; Solarbio, China). These cultured cells were treated with UBCS039 at a concentration of 100 µM for 72 h. This concentration was determined according to our previous published study in which we treated natural killer (NK) cells with UBCS039 [21-23]. Cells that were treated with DMSO and left untreated were used as controls.
Cell proliferation assay
The proliferation of SMMC-7721 cells treated with UBCS039 at a concentration of 100 µM was determined via an RTCA DPlus station (Agilent, USA). Cultured cells were collected in RPMI 1640 medium and added to the E-plate of the equipment for 24 h. Cell proliferation was monitored in real time.
Cell apoptosis assay
The apoptosis of SMMC-7721 cells treated with UBCS039 at a concentration of 100 µM was analyzed via an annexin V-fluorescein isothiocyanate (FITC)/propidine iodide (PI) apoptosis detection kit (Elabscience, USA). Cultured cells were washed and resuspended in 1 × annexin V binding buffer. Annexin V-FITC and PI staining solutions were added. The stained cells were analyzed via a flow cytometer (Apogee A50, NovoCyte D2040R, UK).
Cell migration assay
Cultured SMMC-7721 cells treated with UBCS039 at a concentration of 100 µM were collected and resuspended in serum-free medium. The serum-free cell suspension was added to the upper chamber of a Transwell chamber (Corning, USA), and medium containing 20% FBS was added to the lower chamber. After 24 h of incubation, the cells were fixed with methanol. The cells in the lower chamber were stained with 0.1% crystal violet (Solarbio, China), photographed and counted.
Cellular energy metabolism assay
SMMC-7721 cells were cultured with UBCS039 at a concentration of 100 µM. SMMC-7721 cells at a density of 1 × 104 cells/well were plated in an XFp cell culture plate (Agilent, USA) and cultured in a CO2-free cell culture box overnight at 37 °C for 24 h. The culture medium was discarded, and a detection solution (RPMI 1640 medium containing pyruvate, glucose and glutamine) was added. The XFp cell culture plate was placed in a CO2-free cell culture box at 37 °C for 1 h. A test solution containing rotenone (Rot, 5 µM)/antimycin A (AA, 5 µM) (Agilent, USA) was added to the probe plate for dosing well A, and a test solution containing oligomycin (Agilent, USA) was added to well B. After the probe plate was calibrated in an Agilent Seahorse XFp system, the energy metabolism of the treated cells was examined.
Real-time quantitative PCR
Total RNA was extracted from cultured tumor cells via RNAiso Plus (Takara, Japan). The isolated RNA was reverse transcribed into complementary deoxyribonucleic acid (cDNA) via HiScript III RT SuperMix (Vazyme, China). The cDNA was added to ChamQ Universal SYBR qPCR Master Mix (Vazyme, China), and the mRNA expression of the target genes was examined via fluorescence-based real-time quantitative PCR via a LightCycle96 (Switzerland). Relative mRNA expression was analyzed via the 2-ΔΔ CT method. β-actin mRNA was used as an internal control to quantify the expression levels of target genes. The sequences of the primers used are listed here (Supplementary Material 1, wjon.elmerpub.com).
Isolation of mononuclear cells (MNCs)
Peripheral blood was collected from healthy volunteers (n = 30) to isolate cells for coculture experiments. Ficoll cell separation solution (TBD, China) was added to the diluted blood. After centrifugation, the MNCs were extracted via a routine method, and 10 samples were randomly mixed to form a testing unit. All volunteers provided informed written consent. The study was approved by the Ethics Committee of the Affiliated Hospital of Qingdao University (QYFYWZLL26623) and conducted in compliance with the ethical standards of the responsible institution on human subjects, as well as with the Helsinki Declaration.
Enrichment of naive CD4+ T cells
MNCs were resuspended in phosphate-buffered saline (PBS) containing 2 mM EDTA and 0.5% bovine serum albumin (BSA). Naive CD4+ T-cell Biotin-Antibody Cocktail II (Miltenyi Biotec, Germany), which includes biotinylated monoclonal antibodies against CD8, CD14, CD16, CD19, CD36, CD45RO, CD56, CD123, TCR γ/δ and glycophorin A, was added to the isolated MNCs. Microbeads conjugated to an anti-biotin monoclonal antibody were also added to the mixture. The cell suspension was applied to an LS column with a MACS column (Miltenyi Biotec). The unlabeled cells, representing enriched naive CD4+ T cells, were eluted from the flow-through fraction and collected.
Coculturing tumor cells with CD4+ T cells
A2780, Huh-7, HeLa, MBA-MD-231, MCF-7, SMMC-7721 and SW480 tumor cells were treated with UBCS039 at a final concentration of 100 µM for 72 h. UBCS039 was removed by centrifugation, and the cells were diluted with fresh culture medium. The washing process was performed three times. The treated tumor cells were plated in the upper chamber of a Transwell device with a 0.4-µm pore size (Corning Incorporated, Costar, USA) at a concentration of 3 × 104/100 µL; meanwhile, CD4+ T cells (3 × 104/600 µL) were plated in the lower chamber of the Transwell device. The cells were cocultured for 48 h. The system did not allow direct contact between the tumor cells and the T cells.
Detection of the proportion of Tregs in the coculture system
CD4+ T cells in the lower chamber of a Transwell device were collected and resuspended in PBS. The proportions of Tregs to CD4+ T cells were detected with a Human Regulatory T-cell Staining Kit (Multi Sciences, China). FITC-conjugated anti-CD4 antibody and allophycocyanin (APC)-conjugated anti-CD25 antibody were added to the cell samples, which were then incubated for 15 min in the dark. After the supernatant was removed via centrifugation, the remaining cell pellet was fixed and permeabilized. PE-conjugated anti-FoxP3 antibody was added, and the cells were incubated in the dark at room temperature for 1 h. The proportion of Tregs (CD4+CD25+FoxP3+) in CD4+ T cells was detected via flow cytometry.
Detection of T helper 17 (Th17) cells in the coculture system
CD4+ T cells in the lower chamber of a Transwell device were collected and resuspended in PBS. Th17 cells were analyzed via a Human Th17 Staining Kit (Multi Sciences). PMA/ionomycin and BFA/monensin mixtures were added to the cell samples. The mixture was incubated at 37 °C for 4 h. After centrifugation and resuspension, anti-human FITC-conjugated anti-CD3 antibodies and anti-human CD8α and PerCP-Cy5.5 antibodies were added to the mixture, which was subsequently incubated. Following treatment with FIX & PERM, anti-human PE-IL-17A antibodies were added to the mixture. The frequencies of Th17 cells among CD3+CD8- T cells were measured via flow cytometry.
Detection of cytokine levels in culture medium
Culture medium was collected from cocultures of tumor cell lines and CD4+ T cells. Cytokine levels were measured via a 13-plex Human Inflammation Panel (BioLegend, USA). The cytokine concentrations were determined by flow cytometry (Apogee A50, NovoCyte D2040R, UK). Legend_plex_v8.0 software (BioLegend, USA) was used to analyze the concentrations of interferon (IFN-α2), IFN-γ, interleukin (IL)-1β, IL-6, IL-8, IL-10, IL-12p70, IL-17A, IL-18, IL-23, IL-33, monocyte chemoattractant protein-1 (MCP-1) and tumor necrosis factor (TNF)-α.
ADO assay
Culture medium was collected from cocultures of tumor cells and CD4+ T cells. ADO levels were measured by using an Adenosine Assay Kit (fluorometric) (Abcam, USA). The fluorescence was immediately measured via a microplate reader (Infinite M Plex, Tecan, Switzerland).
Transcriptome analysis of UBCS039-pretreated SMMC-7721 cells
SMMC-7721 cells were pretreated with UBCS039 or DMSO or left untreated. The treated cells were cultured with CD4+ T cells in Transwell as described above. Total RNA was extracted from the cultured SMMC-7721 cells and reverse transcribed into cDNA. The sequencing was performed on an Illumina platform. HISAT2 was used to align the sequences of the clean reads with the specified reference genome to characterize the sequences in the samples. The protein-coding gene expression abundance in each sample was determined via sequence similarity comparison. The number of reads aligned to protein-coding genes in each sample was obtained via Htseq-count software. DESeq2 software was used to standardize the sample gene counts. The base mean value was used to estimate the gene expression level, and the fold change (FC) in expression was calculated. A negative binomial (NB) distribution test was performed to assess the significance of differences in read number. Finally, the differentially expressed protein-coding genes were identified on the basis of the expression FC and significance of the test results. Genes with FC > 1.5 and P < 0.05 were considered differentially expressed. Gene ontology (GO) enrichment analysis was performed on the differentially expressed genes (DEGs) to determine their functions. Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis was performed to identify the pathways in which more than two DEGs were enriched. The results were sorted based on the -log10 P value of each entry and reported from the largest to the smallest value.
Statistical analysis
The LSD method was used for comparisons of two groups, one-way analysis of variance was used to assess the significance of differences among multiple groups, and Pearson’s correlation method was used for correlation analysis. Differences were considered statistically significant when the test standard was α = 0.05 and P < 0.05.
| Results | ▴Top |
UBCS039 suppressed tumor cell proliferation and stimulated apoptosis
SMMC-7721 cells were cultured in 100 µM UBCS039 for 72 h. Because UBCS039 was dissolved in DMSO, DMSO-pretreated SMMC-7721 cells were used as controls. A cell proliferation assay revealed that, compared with DMSO treatment, UBCS039 treatment significantly inhibited the proliferation of SMMC-7721 cells. A cell apoptosis assay revealed that UBCS039 treatment stimulated tumor cell apoptosis (Supplementary Material 2A, B, wjon.elmerpub.com). Transwell assays did not detect a significant difference in cell migration between UBCS039-treated cells and DMSO-treated control cells (Supplementary Material 2C, wjon.elmerpub.com). Metabolic analysis with the Agilent Seahorse XFp system revealed a significantly elevated oxygen consumption rate (OCR) and decreased extracellular acidification rate (ECAR) in SMMC-7721 cells treated with UBCS039, compared with those in cells treated with DMSO (Supplementary Material 2D, wjon.elmerpub.com). Additionally, PCR demonstrated that UBCS039 treatment increased the mRNA expression level of Sirt6 but not that of Sirt1 or Sirt3 in the treated tumor cells (Supplementary Material 3, wjon.elmerpub.com). The above measurements revealed that UBCS039 treatment specifically increased Sirt6 expression rather than that of other Sirt family members and suppressed tumor cell growth.
UBCS039-pretreated tumor cells altered the tumor immune microenvironment
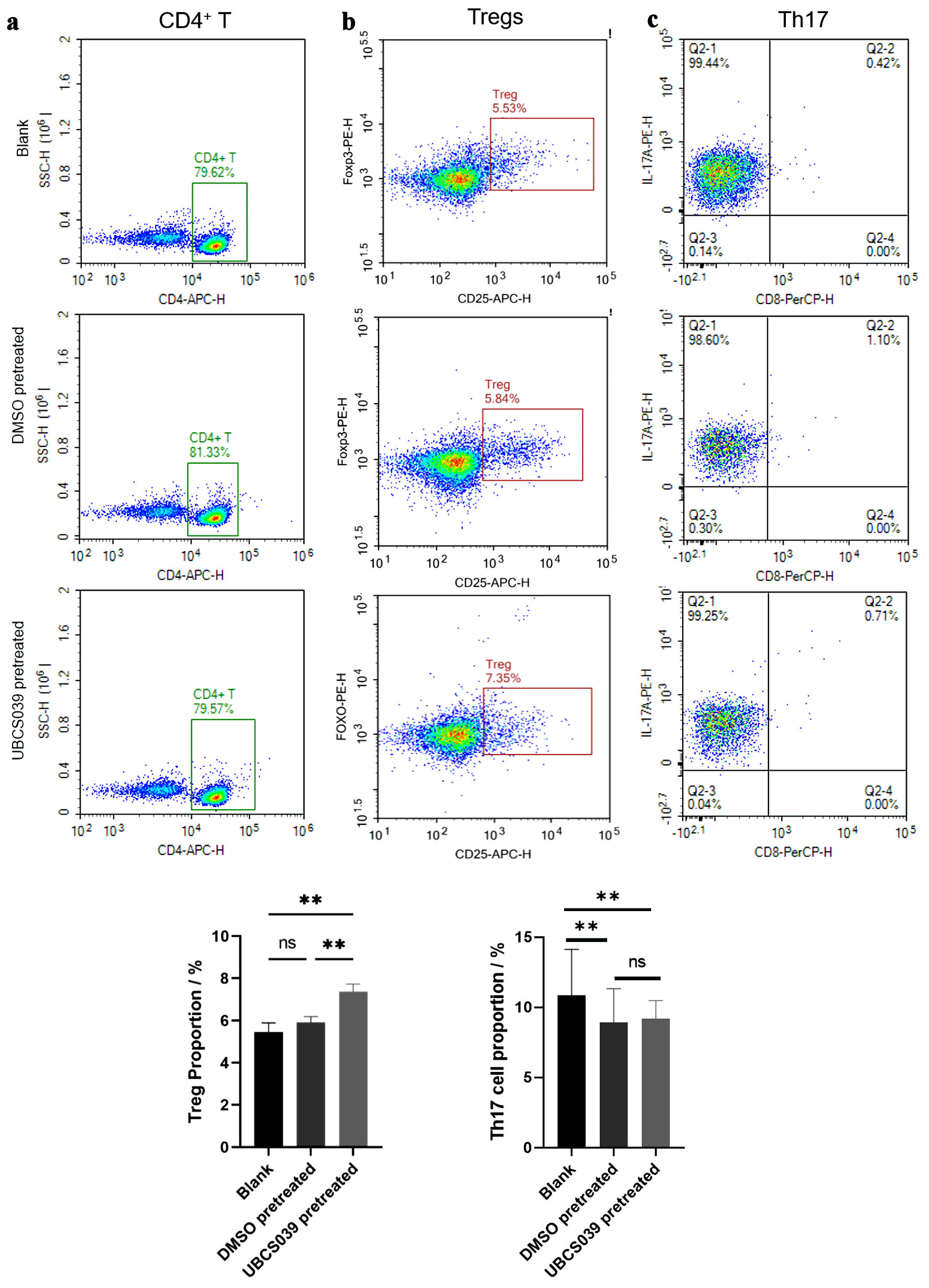
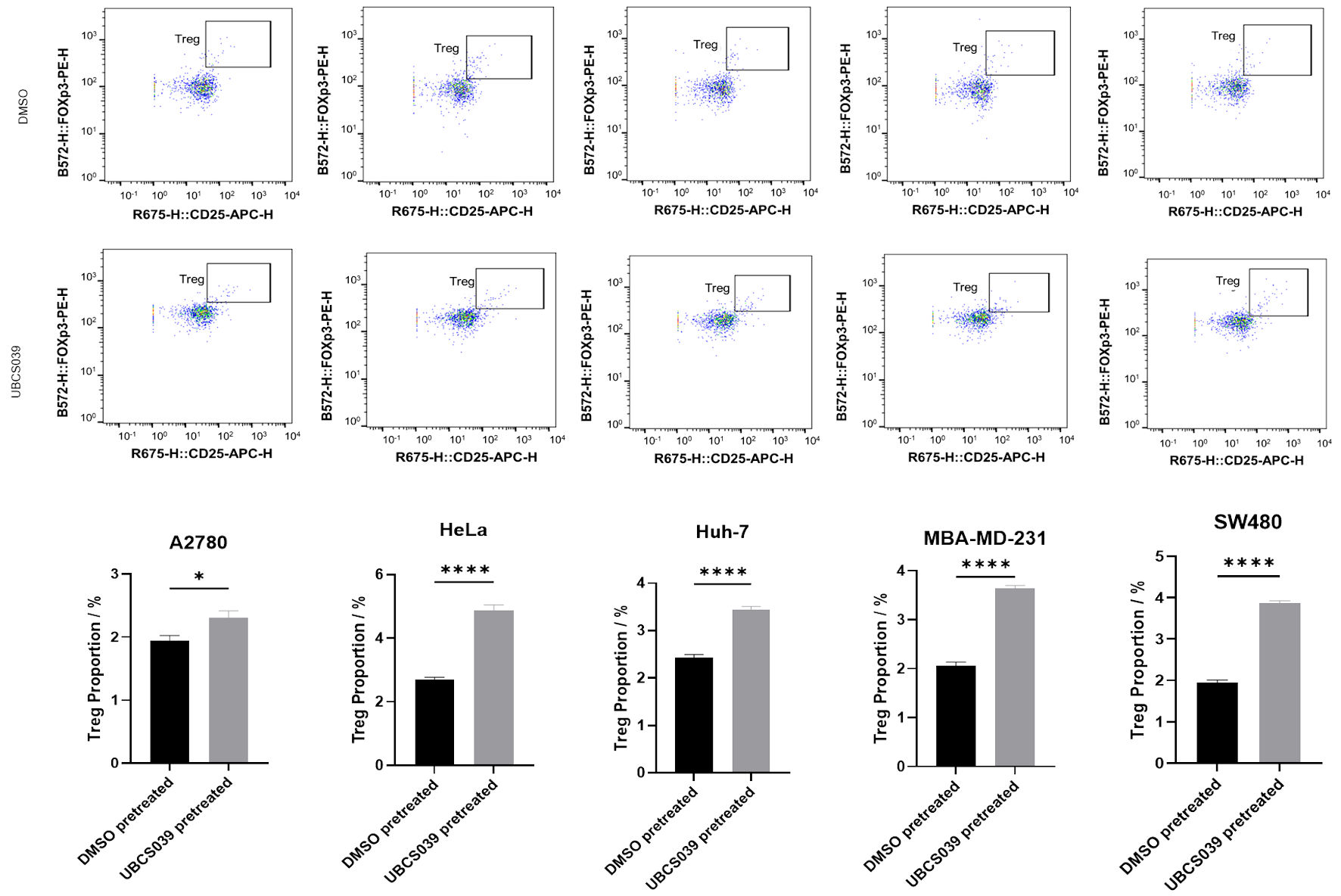
To investigate whether tumor cells with high Sirt6 activity affect CD4+ T-cell differentiation, SMMC-7721 cells were incubated with UBCS039 at a final concentration of 100 µM for 48 h. The tumor cells were washed with PBS after treatment to remove UBCS039 residue and then cultured with human naive CD4+ T cells in Transwell systems. Because UBCS039 was dissolved in DMSO, untreated SMMC-7721 cells and DMSO-pretreated SMMC-7721 cells were used as controls. A significant increase in the Treg (CD25+FoxP3+) proportion was detected in CD4+ T cells following coculture with UBCS039-pretreated SMMC-7721 cells, compared with that in CD4+ T cells following coculture with SMMC-7721 cells without treatment or with DMSO pretreatment; moreover, the proportions of Th17 cells among CD3+CD8- T cells did not differ between those T cells cocultured with DMSO-pretreated SMMC-7721 cells and those T cells cocultured with UBCS039-pretreated SMMC-7721 cells (Fig. 1). To verify the above results, A2780, HeLa, Huh7, MBA-MD-231 and SW480 tumor cells were also treated with 100 µM UBCS039 for 48 h and subsequently cultured with CD4+ T cells in a Transwell after being washed with PBS. Following coculture, the Treg proportions were significantly elevated in CD4+ T cells following coculture with UBCS039-treated tumor cells (Fig. 2). The above results indicate that tumor cells pretreated with UBCS039 can stimulate CD4+ T-cell differentiation into Tregs in the microenvironment.
 Click for large image |
Figure 1. The effect of UBCS039-pretreated SMMC-7721 cells on CD4+ T-cell differentiation. SMMC-7721 cells were treated with UBCS039 or DMSO or left untreated. The proportions of CD4+ T cells (a), Tregs (b), and Th17 cells (c) among the cocultured cells were examined via flow cytometry. The proportions of Tregs (CD4+CD25+FoxP3+) were significantly elevated in CD4+ T cells following coculture with UBCS039-pretreated SMMC-7721 cells. **P < 0.01. Tregs: regulatory T cells; ns: not significant; DMSO: dimethyl sulfoxide; Th17: T helper 17. |
 Click for large image |
Figure 2. Effects of UBCS039-pretreated A2780, HeLa, Huh7, MBA-MD-231 and SW480 cells on CD4+ T-cell differentiation. These tumor cells were treated with UBCS039 or DMSO. The proportions of Tregs among the cocultured cells were examined via flow cytometry. The proportions of Tregs in CD4+ T cells were significantly elevated following coculture with UBCS039-pretreated tumor cells. *P < 0.05, ****P < 0.0001. Tregs: regulatory T cells; ns: not significant; DMSO: dimethyl sulfoxide. |
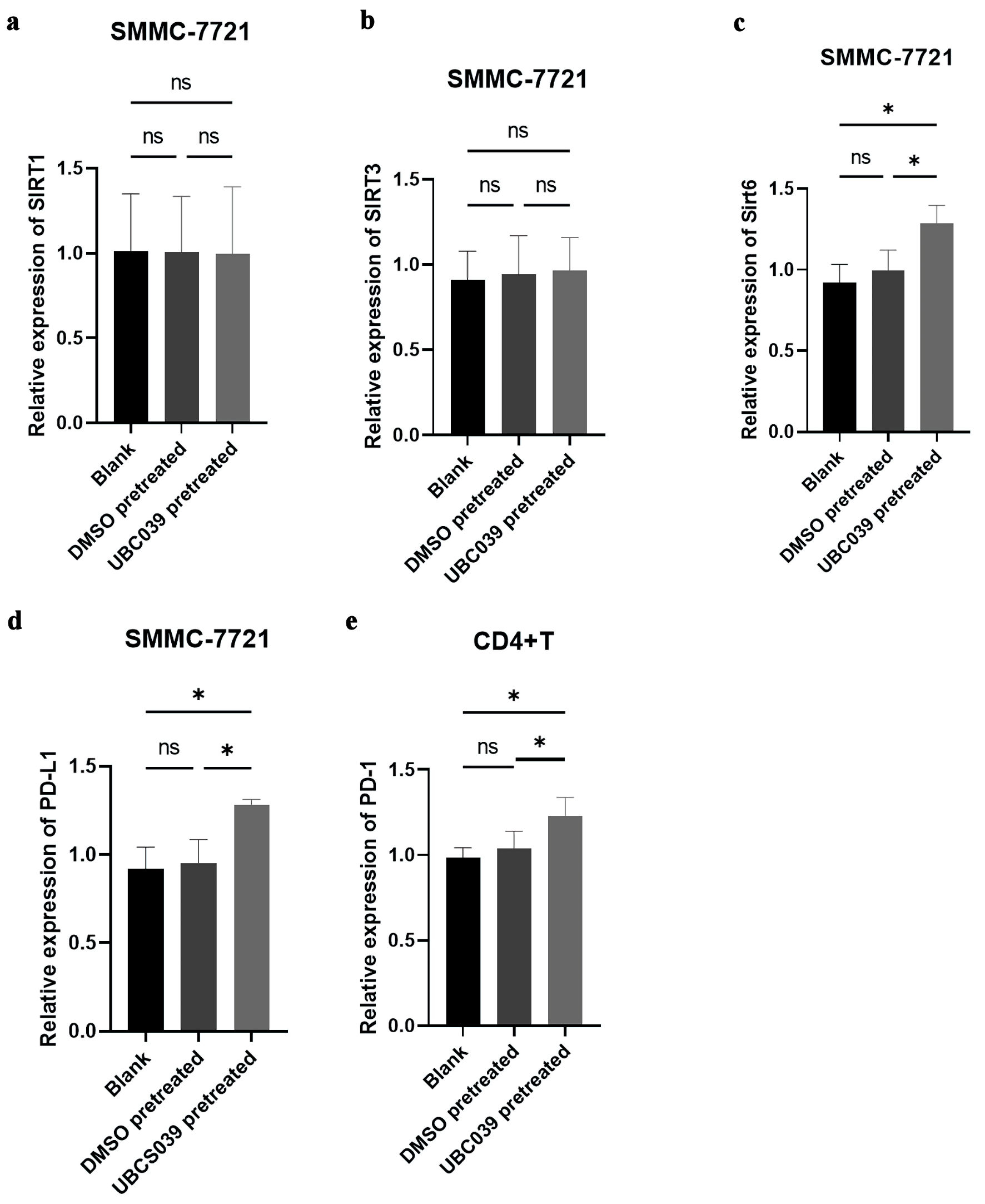
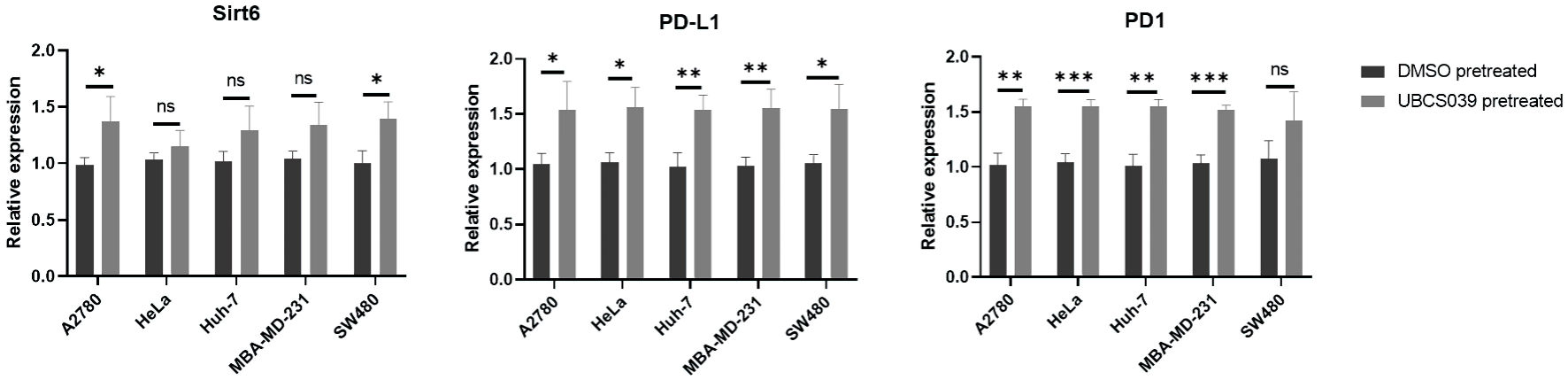
To further understand how UBCS039 treatment affects the immune microenvironment, cocultured SMMC-7721 cells and CD4+ T cells were examined via real-time PCR. The mRNA levels of Sirt1 and Sirt3 did not significantly change in SMMC-7721 cells, regardless of whether the cells were treated with UBCS039 or DMSO or left untreated, but the mRNA levels of Sirt6 and PD-L1 significantly increased in the UBS039-pretreated SMMC-7721 cells. Moreover, the mRNA level of PD-1 was significantly greater in CD4+ T cells following coculture with UBS039-pretreated SMMC-7721 cells than in those cocultured with DMSO-treated tumor cells or untreated tumor cells (Fig. 3). An increased level of Sirt6 was detected in other tumor cell lines following UBCS039 treatment; increased levels of PD-L1 were detected in A2780, HeLa, Huh7 and MBA-MD-231 tumor cell lines; and increased PD-1 levels were detected in CD4+ T cells cocultured with UBCS039-treated A2780, HeLa, Huh7, MBA-MD-231 or SW480 cell lines (Fig. 4). Additionally, flow cytometry revealed a significant decrease in the concentrations of IFN-α2, IFN-γ, IL-10, MCP-1 and TNF-α in culture medium upon coculture of CD4+ T cells and UBS039-pretreated SMMC-7721 cells, compared with those observed upon coculture of CD4+ T cells and DMSO-pretreated SMMC-7721 cells or untreated SMMC-7721 cells. The levels of other cytokines in the culture medium did not change. Furthermore, enzyme-linked immunosorbent assay (ELISA) revealed a significant increase in the ADO level in the culture medium of CD4+ T cells cultured with UBS039-pretreated tumor cells. ADO levels did not significantly change in the coculture medium of CD4+ T cells and DMSO-pretreated SMMC-7721 cells or untreated SMMC-7721 cells (Fig. 5). Significant decreases in the concentrations of IFN-α2, IFN-γ, IL-10 and MCP-1 were also detected in the culture media of cocultured CD4+ T cells and UBS039-pretreated A2780, HeLa, Huh7, MBA-MD-231 or SW480 cells, compared with those in the cocultures of DMSO-pretreated tumor cells and CD4+ T cells, and an increase in the concentration of ADO was detected in the coculture media of UBCS039-pretreated A2780, HeLa and Huh-7 cells and CD4+ T cells (Fig. 6). The above experiments confirmed the increased levels of Sirt6, Tregs, ADO, PD-1 and PD-L1 and decreased levels of the cytokines IFN-α2, IFN-γ, IL-10 and MCP-1 in cocultured CD4+ T cells and UBCS039-treated tumor cells as well as their culture medium.
 Click for large image |
Figure 3. Expression of several key genes in cocultured UBCS039-pretreated SMMC-7721 cells and CD4+ T cells. The mRNA levels of (a) Sirt1, (b) Sirt3, (c) Sirt6, and (d) PD-L1 in SMMC-7721 cells and the mRNA level of (e) PD-1 in CD4+ T cells were determined via real-time PCR. Sirt6 and PD-L1 expression was significantly increased in SMMC-7721 cells treated with UBCS039, and PD-1 expression was significantly increased in CD4+ T cells cocultured with UBCS039-pretreated SMMC-7721 cells. *P < 0.05. Sirt: sirtuin; PD-1: programmed cell death protein-1; PD-L1: programmed cell death-ligand 1; PCR: polymerase chain reaction; ns: not significant. |
 Click for large image |
Figure 4. Expression of several key genes in cocultured UBCS039-pretreated A2780, HeLa, Huh7, MBA-MD-231 and SW480 cells and CD4+ T cells. The mRNA levels of Sirt6 (a) and PD-L1 (b) in tumor cells and the mRNA level of PD-1 (c) in CD4+ T cells were determined via real-time PCR. Sirt6 and PD-L1 expressions were significantly increased in tumor cells pretreated with UBCS039, and PD-1 expression was significantly increased in CD4+ T cells cocultured with UBCS039-pretreated tumor cells. *P < 0.05, **P < 0.01, and ***P < 0.001. Sirt: sirtuin; PD-1: programmed cell death protein-1; PD-L1: programmed cell death-ligand 1; PCR: polymerase chain reaction; ns: not significant. |
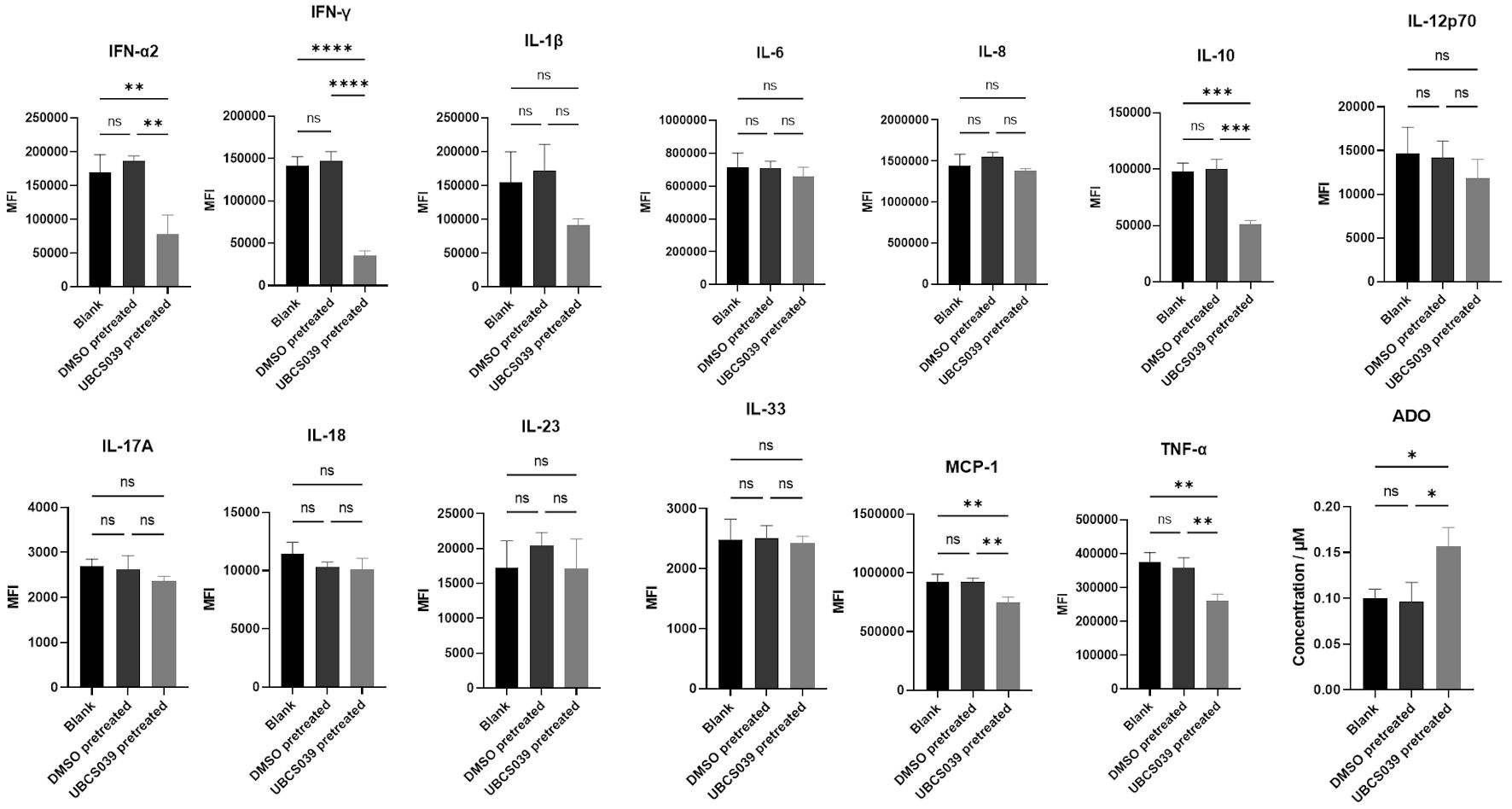 Click for large image |
Figure 5. Cytokine levels in the supernatants of cocultures containing SMMC-7721 cells and CD4+ T cells. SMMC-7721 cells were treated with UBCS039 or DMSO or left untreated. The pretreated SMMC-7721 cells were then cocultured with CD4+ T cells. The culture medium was assessed via flow cytometry. The ADO level in the culture medium was determined via ELISA. IFN-α2, IFN-γ, IL-10, MCP-1 and TNF-α levels were lower in the culture media of cocultures containing UBCS039-pretreated SMMC-7721 cells and CD4+ T cells, and ADO levels were significantly greater in the coculture media. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001. MFI: mean fluorescence intensity; IL: interleukin; IFN: interferon; ADO: adenosine; ELISA: enzyme-linked immunosorbent assay; MCP-1: monocyte chemoattractant protein-1; TNF: tumor necrosis factor; DMSO: dimethyl sulfoxide. |
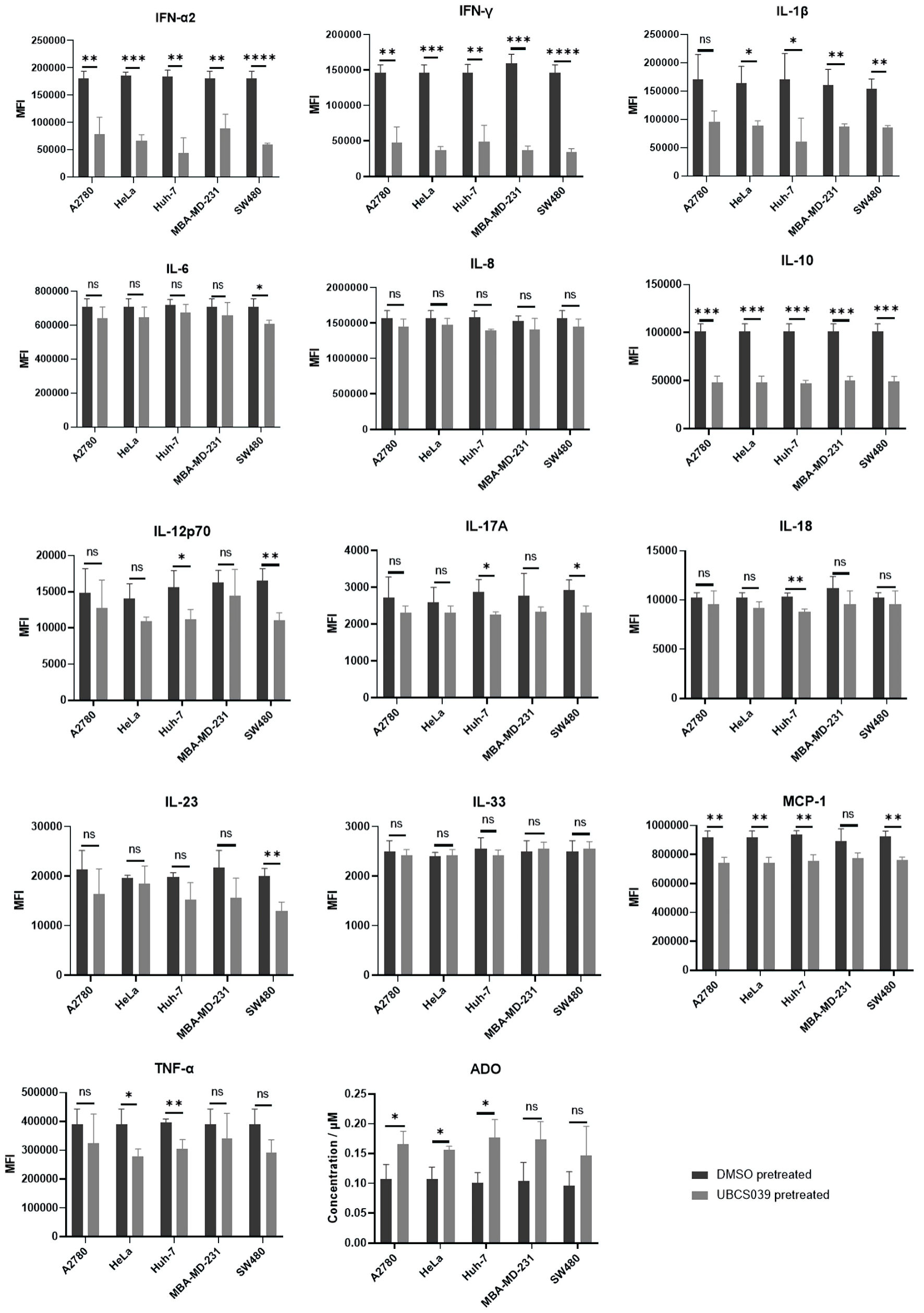 Click for large image |
Figure 6. Cytokine levels in the supernatants of cocultures containing CD4+ T cells and A2780, HeLa, Huh7, MBA-MD-231 or SW480 cells. These tumor cells were pretreated with UBCS039 or DMSO. The pretreated tumor cells were then cocultured with CD4+ T cells. The culture medium was assessed via flow cytometry. The ADO level in the culture medium was determined via ELISA. The levels of IFN-α2, IFN-γ and IL-10 were lower in the culture media of cocultures containing UBCS039-pretreated tumor cells and CD4+ T cells, and the ADO levels were significantly greater in the coculture media of CD4+ T cells and UBCS039-pretreated A2780, HeLa and Huh-7 cells. *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001. MFI: mean fluorescence intensity; IL: interleukin; IFN: interferon; ADO: adenosine; ELISA: enzyme-linked immunosorbent assay; MCP-1: monocyte chemoattractant protein-1; TNF: tumor necrosis factor; DMSO: dimethyl sulfoxide. |
UBCS039 changed the expression profiles and regulatory pathways of tumor cells
To understand the regulatory mechanism by which Sirt6 in tumor cells affects the immune microenvironment, UBCS039-pretreated SMMC-7721 cells were cultured with CD4+ T cells, and the pretreated tumor cells were examined via transcriptomic analysis. The expression files of UBCS039-pretreated SMMC-7721 cells, DMSO-pretreated SMMC-7721 cells and untreated SMMC-7721 cells were compared. The top DEGs between the expression files of UBCS039-treated SMMC-7721 cells and SMMC-7721 cells without treatment (Fig. 7a, c) were completely different from those of DMSO-treated SMMC-7721 cells and SMMC-7721 cells without treatment (Fig. 7b, c), indicating that UBCS039 treatment affects specific pathways in tumor cells to affect CD4+ cells. The DEGs, including BASP1, CPS1, FOXA2, GSTP1, GNG11, MFAP5, NNMT, RASEF, SMOC1, SERPINA6 and ZNF844, between the UBCS039-pretreated SMMC-7721 cells and the untreated tumor cells, were also differentially expressed between the UBCS039-pretreated SMMC-7721 cells and the DMSO-treated cells, indicating that the increased Sirt6 activity and expression induced by UBCS039 specifically regulated the expression of these genes.
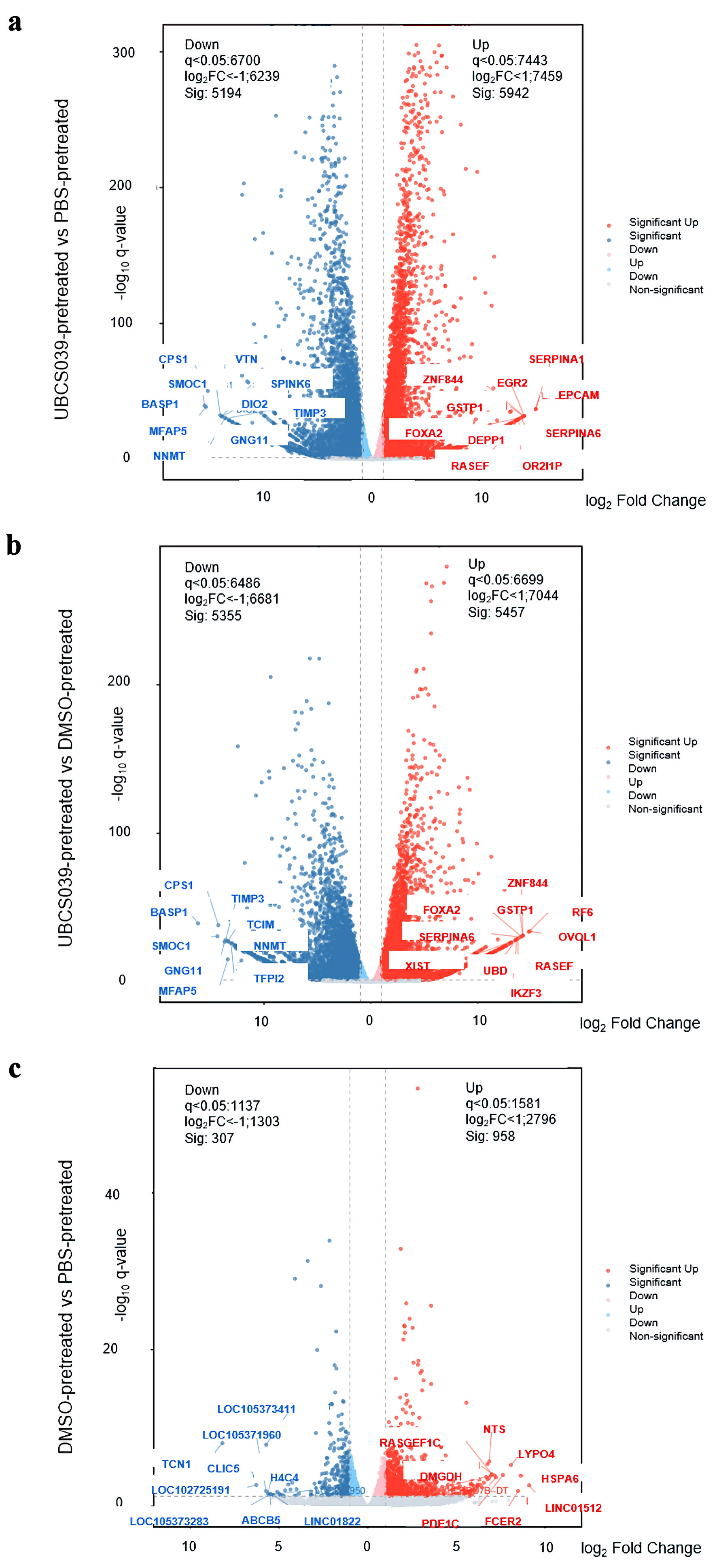 Click for large image |
Figure 7. Transcriptome analysis of UBCS039-pretreated SMMC-7721 cells. Volcano plots were generated by plotting the log (base 2) of the fold change (FC) and P values of the t test results (base 10) between the UBCS039-pretreated SMMC-7721 cells and the untreated SMMC-7721 cells (a), UBCS039-pretreated SMMC-7721 cells and DMSO-pretreated SMMC-7721 cells (b), and DMSO-pretreated SMMC-7721 cells and SMMC-7721 cells without treatment (c). The analysis revealed significant changes in the gene expression profile of SMMC-7721 cells following UBCS039 treatment. DMSO: dimethyl sulfoxide. |
The specific expression of these DEGs was verified via real-time PCR in pretreated SMMC-7721 cells, in which those cells treated with DMSO or left untreated were used as controls. The mRNA levels of BASP1, CPS1, GNG11, MFAP5, NNMT and SMOC1 were significantly lower, and the levels of FOXA2, GSTP1, RASEF and ZNF844 were significantly greater in SMMC-7721 cells following UBCS039 treatment, which was in accordance with the results of the transcriptomic analysis, except for those of SERPINA6 (Supplementary Material 4, wjon.elmerpub.com). The decreased expression of BASP1, CPS1, GNG11, MFAP5, NNMT and SMOC1 and the increased expression of FOXA2, GSTP1, RASEF and ZNF844, but not SERPINA6, were also detected in A2780, HeLa, Huh7, MBA-MD-231 and SW480 tumor cells that were pretreated with UBCS039 and cocultured with CD4+ T cells (Supplementary Material 5, wjon.elmerpub.com). These results indicate that Sirt6 mainly regulates the expression of BASP1, CPS1, GNG11, MFAP5, NNMT, SMOC1, FOXA2, GSTP1, RASEF and ZNF844 in tumor cells.
KEGG enrichment analysis was performed on the DEGs. The expression profiles of UBCS039-pretreated SMMC-7721 cells were compared with those of DMSO-pretreated SMMC-7721 cells and SMMC-7721 cells without treatment. Pathways associated with activated genes, including adherens junction, TNF signaling, circadian rhythm, glucagon signaling, parathyroid hormone synthesis and neutrophil extracellular trap formation, were enriched by comparing the expression data of UBCS039-pretreated SMMC-7721 cells and those of untreated cells. These pathways were also enriched when the expression data of UBCS039-pretreated SMMC-7721 cells and DMSO-treated cells were compared (Fig. 8), indicating that Sirt6 specifically regulates these pathways in treated tumor cells to affect CD4+ T-cell differentiation, as well as other immune surveillance-related factors in the microenvironment.
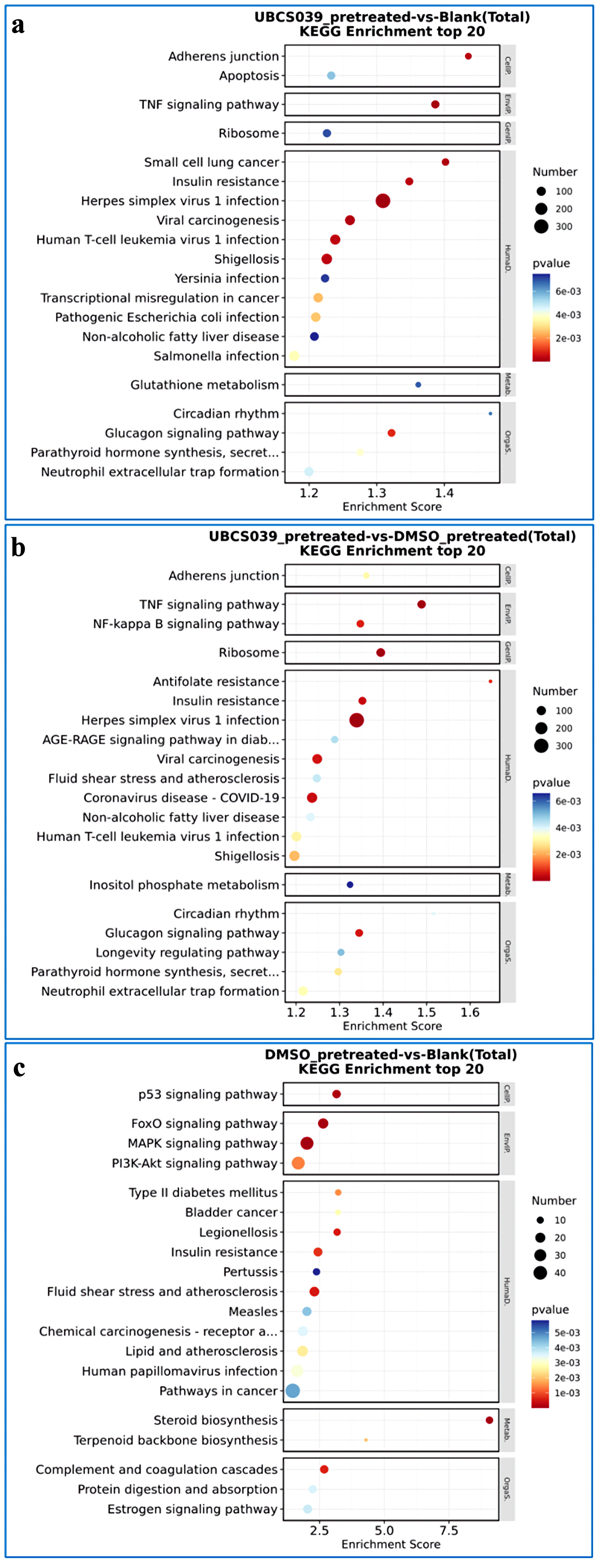 Click for large image |
Figure 8. KEGG enrichment analysis of UBCS039-pretreated SMMC-7721 cells. KEGG enrichment analysis was performed based on DEGs between UBCS039-pretreated SMMC-7721 cells and untreated SMMC-7721 cells (a), UBCS039-pretreated SMMC-7721 cells and DMSO-pretreated SMMC-7721 cells (b), and DMSO-pretreated SMMC-7721 cells and untreated SMMC-7721 cells (c). The analysis revealed some significant changes in regulatory pathways in SMMC-7721 cells following UBCS039 treatment. KEGG: Kyoto Encyclopedia of Genes and Genomes; DEGs: differentially expressed genes; DMSO: dimethyl sulfoxide. |
| Discussion | ▴Top |
Increased numbers of Tregs are observed in the peripheral circulation and tumor tissues of many cancer patients [24, 25]. High numbers of infiltrating Tregs are strongly correlated with shorter overall survival in cancer patients [26]. However, the cellular mechanisms underlying the activation or expansion of Tregs in the TME by tumor cells have largely not been explored. In the present study, an increase in the Treg population was detected in CD4+ T cells cocultured with UBSC039-pretreated tumor cells, including A2780, HeLa, Huh7, MBA-MD-231, SMMC-7721 and SW480 cells. We also detected increased Sirt6 expression in these UBCS039-treated cells. These observations suggest that tumor cells with high Sirt6 activity and expression can stimulate more CD4+ T-cell differentiation into Tregs in the TME.
This study revealed significant reductions in the IFN-α2, IL-10, IFN-γ and MCP-1 levels in the coculture medium of UBCS039-treated tumor cells and CD4+ T cells. The alteration in the production of these cytokines may be due to the interaction between UBCS039-pretreated tumor cells and CD4+ T cells in the microenvironment. The tumor expression levels of chemokine (C-C motif) ligand (CCL)2 (MCP-1), CCL22, and chemokine (C-X-C motif) ligand (CXCL)12 are correlated with the intratumoral expression of the Treg markers FOXP3, CD33 and NCF2 [27]. IFN-γ production has been reported to inhibit Treg-cell differentiation [28, 29] and limit CD25 and Foxp3 expression in CD4+Foxp3+ Tregs [30]. One study revealed that CD4+CD25-Foxp3+ Tregs were generated when naive CD4+ T cells were cocultured with either adipose-derived stem cells (ASCs) or their related supernatant, while IFN-γ was downregulated [31]. Another study revealed that transforming growth factor (TGF)-β1-overexpressing mesenchymal stem cells also regulate Treg differentiation by downregulating IFN-γ production [32]. Additionally, a study reported that the level of IFN-γ was greater in Sirt5 knockout mice than in control mice, suggesting that the Sirt family may downregulate IFN-γ production [33]. IFN-γ functions through the suppression of Tregs [34]. Several studies have reported that Sirt6 changes the expression of proinflammatory cytokines and chemokines [35]. The findings of others supported our findings with UBCS039-treated tumor cells. Thus, we suggest that tumor cells with high Sirt6 activity and expression stimulate CD4+ T-cell differentiation into Tregs to interrupt immune surveillance by reducing IFN-γ production. Tregs secrete IL-10 and TGF-β to contribute to immune surveillance. However, we detected lower IL-10 levels in the culture medium of CD4+ T cells cocultured with UBCS039-pretreated tumor cells than in those cocultured with DMSO-pretreated tumor cells or untreated tumor cells. We repeated this experiment and obtained similar results. Most likely, the pretreatment of tumor cells with UBCS039 suppressed IL-10 production. This phenomenon needs further investigation. Under in vivo conditions, many immune cells participate in the composition of TME, in addition to Tregs and tumor cells.
This study revealed increased PD-L1 expression in UBCS039-pretreated tumor cells and PD-1 expression in cocultured CD4+ T cells. It is well known that the PD-1/PD-L1 axis can influence Treg differentiation beyond regulating cell exhaustion [36-38]. The PD-1/PD-L1 pathway is critical for promoting Treg cell development and inhibiting the Th17 response during pregnancy [39]. Bispecific anti-PD-(L)1/TGF-β inhibitors hamper Treg expansion [40]. PD-L1 Fc, an activator of the PD-1/PD-L1 pathway, increased the percentage of Treg cells and increased Foxp3 mRNA expression [40, 41]. Blocking the PD-1/PD-L1 pathway may suppress the differentiation of Treg cells [42]. PD-1 also activated FoxP3+ Tregs [41, 42]. Although there are no reports about the regulatory effect of Sirt6 on PD-1/PD-L1, a study showed that PD-L1 can promote the growth and metastasis of cervical cancer by activating the ITGB4/SNAI1/SIRT3 signaling pathway [43]. Estradiol also facilitates the growth and metastasis of non-small cell lung cancer in nude mice via the SIRT1/FOXO3a/PD-L1 axis [44]. Thus, increased Sirt6 activity in tumor cells likely stimulates CD4+ T-cell differentiation into Tregs by partially activating the PD-1/PD-L1 axis in the microenvironment to impede immune surveillance.
This study revealed an increase in the level of ADO in the coculture medium of CD4+ T cells and UBCS039-pretreated tumor cells. It has been confirmed that extracellular ADO can lead to strong immunosuppressive effects, facilitating tumor escape through the inhibition of CD4+, CD8+, NK and dendritic cells and the stimulation of Tregs [45]. A subset of inducible Tregs expressing the ectonucleotidases CD39 and CD73 can hydrolyze ATP to 5'-AMP and ADO and thus mediate immune cells that express ADO receptors [46, 47]. Thus, it is reasonable that coculturing CD4+ T cells and tumor cells with high Sirt6 activity produces more ADO to impede immune surveillance.
A transcriptomic analysis of SMMC-7721 cell tumors was conducted following UBCS039 treatment, and the common DEGs were detected by comparing the expression profiles of UBCS039-treated, DMSO-treated and untreated SMMC-7721 cells. Real-time PCR confirmed the decreased mRNA expression of BASP1, CPS1, GNG11, MFAP5, NNMT and SMOC1 and increased mRNA expression of FOXA2, GSTP1, RASEF and ZNF844 in UBCS039-pretreated A2780, HeLa, Huh7, MBA-MD-231, SMMC-7721 and SW480 tumor cells that were cocultured with CD4+ T cells. BASP1 expression is decreased in many tumor tissues and human tumor cell lines [48-51]. Restoration of BASP1 expression results in extensive antitumor activity [48, 51, 52]. Nuclear BASP1 converts the Wilms tumor transcription factor WT1 from an oncoprotein into a tumor suppressor and significantly suppresses cell proliferation, migration and invasion; promotes apoptosis; and suppresses v-myc-induced cell transformation [53, 54]. CPS1 expression is downregulated in hepatocellular carcinoma [55, 56]. Inhibition of CPS1 upregulated cyclin A2 and cyclin D1, two cell cycle regulatory proteins, by stabilizing the oncoprotein c-Myc at the posttranscriptional level [55]; GNG11 was downregulated in ovarian cancer patients [57]; MFAP5 induced apoptosis and inhibited migration and invasion, but this process was downregulated in the tumor stroma in cervical cancer [58-60]; SMOC1 suppressed proliferation, colony formation and in vivo tumor formation; and SMOC1 expression was also decreased in tumor tissues [61]. These studies support that BASP1, CPS1, GNG11, MFAP5 and SMOC1 are antitumor genes and that their expression is decreased in tumor tissues. On the other hand, FOXA2 expression is significantly increased in esophageal squamous cell carcinoma, prostate cancer, colorectal cancer, and esophageal cancer [62-64]; elevated RASEF expression promotes cell growth via ERK signaling, and this expression is associated with poor prognosis in non-small cell lung cancer [65]; and GSTP1 methylation has been detected in several malignant tumors, such as prostate, breast, lung and hepatocellular carcinoma [66, 67]. These studies support that FOXA2, GSTP1 and RASEF are oncogenes that are highly expressed in tumor tissues. Additionally, some reports confirmed the regulatory effect of Sirt6 on the expression of these top DEGs. For example, Sirt6 can interact directly with FOXA2 in vivo and in vitro and plays a critical role in hepatocellular carcinoma progression [68]. The findings of the above studies corresponded with our findings. Our study suggested that increasing Sirt6 activity and expression downregulates BASP1, CPS1, GNG11, MFAP5 and SMOC1 expression and upregulates FOXA2, GSTP1 and RASEF expression to promote tumorigenesis. However, this pathway warrants further investigation.
Many studies have also reported that BASP1, SMOC1 and ZNF844 are involved in the tumor immune microenvironment. BASP1 expression is positively correlated with the expression of immune checkpoints and immune cell markers, as well as with immune cell infiltration [69]. Patients with high BASP1 expression exhibited a better response to anti-PD-1 immunotherapy and a greater response to immune checkpoint inhibitors. BASP1 was also shown to be positively correlated with T-cell dysfunction and immune escape [70]. SMOC1 expression was negatively correlated with the levels of infiltrating B cells, CD8+ T cells, CD4+ T cells, macrophages, neutrophils and dendritic cells [71]; ZNF844 expression was moderately inversely correlated with helper T-cell subtype 1 infiltration [72]. Thus, increased Sirt6 activity and expression in tumor cells may downregulate BASP1 and SMOC1 expression and upregulate ZNF844 expression to change the tumor immune microenvironment to favor tumor growth. Additionally, a study showed that GSTP1 prevents LPS-induced TNF-α, IL-1β and MCP-1 production [73]. Thus, it is highly possible that increased Sirt6 activity and expression following UBCS039 treatment decreased IL-1β and MCP-1 production by upregulating GSTP1 expression in the present study, although the mechanism is unknown.
The activation of several pathways, such as the adherens junction, TNF signaling, and circadian rhythm pathways, was detected via KEGG enrichment analysis in SMMC-7721 cells following UBCS039 treatment. It has been reported that blocking the TNF pathway in a variety of tumor models inhibits tumor growth [74]. Moreover, Sirt6 knockdown increased the mRNA levels of NF-κB-associated TNF-α signaling factors [75, 76]. Additionally, E-cadherin, a cell adhesion protein, accumulates in tumor cell adherens junctions and is less stable than it is in normal epithelial cells. E-cadherin-based adherens junctions are essential for the migration and invasion of carcinoma cells. Downregulation of E-cadherin expression is considered a poor prognostic factor for patients with tumors [77]. Sirt6 has been demonstrated to be a critical regulator of circadian transcription that serves as an interface with metabolic homeostasis [78]. Thus, increased Sirt6 activity and expression in tumor cells likely activate adherens junctions, TNF signaling and the circadian rhythm to change the TME to favor tumor cell growth.
In SMMC-7721 cell culture, UBCS039 treatment suppressed cell proliferation and stimulated apoptosis. Furthermore, UBCS039 increased the OCR and reduced the ECAR, thereby reversing the metabolic balance in tumor cells. The increase in aerobic glycolysis in tumor cells is called the Warburg effect [79]. A study suggested that Sirt6 is a negative regulator of the Warburg effect [14]. These in vitro experiments suggest that increased Sirt6 activity inhibits tumor malignancy in vitro. On the other hand, the present study revealed that pretreatment of tumor cells with UBSC039 can increase ADO and PD-1/PD-L1 levels as well as Treg abundance to favor tumor cell immune escape. Thus, UBCS039 treatment has completely different effects on tumor cell growth in the TME in vivo than in vitro. Evidence has indicated dual roles of Sirt6 in tumor onset and progression and that these opposite outcomes occur in cancer [80].
In the present study, Sirt6 mRNA expression increased in UBCS-039-treated tumor cells and in UBCS-039-pretreated tumor cells cocultured with CD4+ T cells, but Sirt1 and Sirt3 expression did not significantly change. This result suggested that UBCS039 specifically enhanced Sirt6 activity and Sirt6 expression rather than Sirt1 and Sirt3 expression. In our study, transcriptomic analysis revealed the downregulation of the antitumor genes BASP1, CPS1, GNG11, MFAP5, NNMT and SMOC1; the upregulation of the tumor-promoting genes FOXA2, GSTP1, RASEF and ZNF844; and the activation of adherens junctions and the TNF-signaling and circadian rhythm pathways in UBCS039-pretreated SMMC-7721 cells. However, it is unclear how Sirt6 regulates key genes and pathways. Most likely, ChIP-seq or CRISPR-based gene editing techniques can be used to investigate how Sirt6 directly regulates key genes.
Conclusions
The present study revealed that human tumor cells with increased Sirt6 activity can stimulate CD4+ T cells to differentiate into Tregs and cooperate with CD4+ T cells to increase ADO production and PD-1/PD-L1 activity, which impedes immune surveillance and consequently favors tumor growth in the TME. It is possible that the tumor cells with high Sirt6 activity play the tumor-promoting role by altering proinflammatory cytokine production, especially by downregulating IFN-γ production. The tumor cells pretreated with UBCS039 act by downregulating BASP1, CPS1, GNG11, MFAP5, NNMT and SMOC1 expression and upregulating FOXA2, GSTP1, RASEF and ZNF844 expression. The above results may be helpful for understanding the tumor immune escape mechanism during tumorigenesis and suggest that Sirt6 is a key gene for immune surveillance.
| Supplementary Material | ▴Top |
Suppl 1. Primer sequences.
Suppl 2. The effect of UBCS039 on SMMC-7721 cell growth.
Suppl 3. The effect of UBCS039 on Sirt gene expression in SMMC-7721 cells.
Suppl 4. Verification of the special expression of DEGs identified via transcriptomic analysis in SMMC-7721 cells via real-time PCR.
Suppl 5. Verification of the special expression of DEGs identified via transcriptomic analysis in A2780, HeLa, Huh7, MBA-MD-231 and SW480 cells via real-time PCR.
Acknowledgments
None to declare.
Financial Disclosure
This work was supported in part by the Shandong Provincial Key R & D Program (2023CXPT040) and Science and Technology Projects in Qingdao West Coast New District (2021-3).
Conflict of Interest
The authors declare no potential conflict of interest.
Informed Consent
All volunteers provided informed written consent.
Author Contributions
NY Zhang and WY Liu performed data curation, formal analysis and investigation. KH Fang performed data curation, formal analysis and investigation, and prepared the original draft. XT Chang designed the study, provided project administration, prepared original draft, and re-edited the manuscript.
Data Availability
The datasets in this study are available from the corresponding author on reasonable request.
| References | ▴Top |
- Roth M, Wang Z, Chen WY. Sirtuins in hematological aging and
malignancy. Crit Rev Oncog. 2013;18(6):531-547.
doi pubmed - Bandopadhyay S, Prasad P, Ray U, Das Ghosh D, Roy SS. SIRT6
promotes mitochondrial fission and subsequent cellular invasion in ovarian cancer. FEBS Open
Bio. 2022;12(9):1657-1676.
doi pubmed - Liu M, Yu J, Jin H, Wang S, Ding J, Xing H, He S, et al.
Bioinformatics analysis of the SIRT family members and assessment of their potential clinical
value. Onco Targets Ther. 2021;14:2635-2649.
doi pubmed - Ding Y, Wu S, Huo Y, Chen X, Chai L, Wang Y, Wang X, et al.
Inhibition of Sirt6 suppresses tumor growth by inducing G1/S phase arrest in renal cancer cells.
Int J Clin Exp Pathol. 2019;12(7):2526-2535.
pubmed - Geng CH, Zhang CL, Zhang JY, Gao P, He M, Li YL.
Overexpression of Sirt6 is a novel biomarker of malignant human colon carcinoma. J Cell
Biochem. 2018;119(5):3957-3967.
doi pubmed - Wu X, Wang S, Zhao X, Lai S, Yuan Z, Zhan Y, Ni K, et al.
Clinicopathological and prognostic value of SIRT6 in patients with solid tumors: a meta-analysis
and TCGA data review. Cancer Cell Int. 2022;22(1):84.
doi pubmed - Zhang Z, Ha SH, Moon YJ, Hussein UK, Song Y, Kim KM, Park
SH, et al. Inhibition of SIRT6 potentiates the anti-tumor effect of doxorubicin through
suppression of the DNA damage repair pathway in osteosarcoma. J Exp Clin Cancer Res.
2020;39(1):247.
doi pubmed - Fu W, Li H, Fu H, Zhao S, Shi W, Sun M, Li Y. The SIRT3 and
SIRT6 Promote Prostate Cancer Progression by Inhibiting Necroptosis-Mediated Innate Immune
Response. J Immunol Res. 2020;2020:8820355.
doi pubmed - Tian J, Yuan L. Sirtuin 6 inhibits colon cancer progression
by modulating PTEN/AKT signaling. Biomed Pharmacother. 2018;106:109-116.
doi pubmed - Zhang C, Yu Y, Huang Q, Tang K. SIRT6 regulates the
proliferation and apoptosis of hepatocellular carcinoma via the ERK1/2 signaling pathway. Mol
Med Rep. 2019;20(2):1575-1582.
doi pubmed - Becherini P, Caffa I, Piacente F, Damonte P, Vellone VG,
Passalacqua M, Benzi A, et al. SIRT6 enhances oxidative phosphorylation in breast cancer and
promotes mammary tumorigenesis in mice. Cancer Metab. 2021;9(1):6.
doi pubmed - Preyat N, Leo O. Sirtuin deacylases: a molecular link
between metabolism and immunity. J Leukoc Biol. 2013;93(5):669-680.
doi pubmed - Pillai VB, Gupta MP. Is nuclear sirtuin SIRT6 a master
regulator of immune function? Am J Physiol Endocrinol Metab.
2021;320(3):E399-E414.
doi pubmed - Parenti MD, Grozio A, Bauer I, Galeno L, Damonte P, Millo E,
Sociali G, et al. Discovery of novel and selective SIRT6 inhibitors. J Med Chem.
2014;57(11):4796-4804.
doi pubmed - Li Y, Jin J, Wang Y. SIRT6 widely regulates aging, immunity,
and cancer. Front Oncol. 2022;12:861334.
doi pubmed - Nishikawa H, Koyama S. Mechanisms of regulatory T cell
infiltration in tumors: implications for innovative immune precision therapies.
J Immunother Cancer. 2021;9(7):e002591.
doi pubmed - Santiago-Sanchez GS, Hodge JW, Fabian KP. Tipping the
scales: Immunotherapeutic strategies that disrupt immunosuppression and promote immune
activation. Front Immunol. 2022;13:993624.
doi pubmed - Malla RR, Vasudevaraju P, Vempati RK, Rakshmitha M, Merchant
N, Nagaraju GP. Regulatory T cells: Their role in triple-negative breast cancer progression and
metastasis. Cancer. 2022;128(6):1171-1183.
doi pubmed - You W, Rotili D, Li TM, Kambach C, Meleshin M, Schutkowski
M, Chua KF, et al. Structural basis of sirtuin 6 activation by synthetic small molecules. Angew
Chem Int Ed Engl. 2017;56(4):1007-1011.
doi pubmed - Iachettini S, Trisciuoglio D, Rotili D, Lucidi A, Salvati E,
Zizza P, Di Leo L, et al. Pharmacological activation of SIRT6 triggers lethal autophagy in human
cancer cells. Cell Death Dis. 2018;9(10):996.
doi pubmed - Wang H, Fang K, Yan W, Chang X. T-Cell Immune Imbalance in
Rheumatoid Arthritis Is Associated with Alterations in NK Cells and NK-Like T Cells Expressing
CD38. J Innate Immun. 2022;14(2):148-166.
doi pubmed - Wang H, Li S, Zhang G, Wu H, Chang X. Potential therapeutic
effects of cyanidin-3-O-glucoside on rheumatoid arthritis by relieving inhibition of CD38+ NK
cells on Treg cell differentiation. Arthritis Res Ther. 2019;21(1):220.
doi pubmed - Zhang X, Wang H, Song X, Song Y, He G, Fang K, Chang X.
Compound 78c exerts a therapeutic effect on collagen-induced arthritis and rheumatoid arthritis.
Clin Exp Rheumatol. 2023;41(7):1384-1395.
doi pubmed - Sarkar T, Dhar S, Sa G. Tumor-infiltrating T-regulatory
cells adapt to altered metabolism to promote tumor-immune escape. Curr Res Immunol.
2021;2:132-141.
doi pubmed - Scott EN, Gocher AM, Workman CJ, Vignali DAA. Regulatory T
cells: barriers of immune infiltration into the tumor microenvironment. Front Immunol.
2021;12:702726.
doi pubmed - Dixon ML, Leavenworth JD, Leavenworth JW. Lineage
reprogramming of effector regulatory T cells in cancer. Front Immunol. 2021;12:717421.
doi pubmed - Su KF, Peng Y, Yu HC. Prognostic value of regulator T cells
in patients with pancreatic cancer: a systematic review and meta-analysis. Eur Rev Med Pharmacol
Sci. 2022;26(8):2906-2917.
doi pubmed - Hariyanto AD, Permata TBM, Gondhowiardjo SA. Role of
CD4(+)CD25(+)FOXP3(+) T(Reg) cells on tumor immunity. Immunol Med. 2022;45(2):94-107.
doi pubmed - Chen BJ, Zhao JW, Zhang DH, Zheng AH, Wu GQ. Immunotherapy
of cancer by targeting regulatory T cells. Int Immunopharmacol. 2022;104:108469.
doi pubmed - Muthuswamy R, Corman JM, Dahl K, Chatta GS, Kalinski P.
Functional reprogramming of human prostate cancer to promote local attraction of effector CD8(+)
T cells. Prostate. 2016;76(12):1095-1105.
doi pubmed - Lee S, Park K, Kim J, Min H, Seong RH. Foxp3 expression in
induced regulatory T cells is stabilized by C/EBP in inflammatory environments. EMBO Rep.
2018;19(12):e45995.
doi pubmed - Olalekan SA, Cao Y, Hamel KM, Finnegan A. B cells expressing
IFN-gamma suppress Treg-cell differentiation and promote autoimmune experimental arthritis.
Eur J Immunol. 2015;45(4):988-998.
doi pubmed - St Rose MC, Taylor RA, Bandyopadhyay S, Qui HZ, Hagymasi AT,
Vella AT, Adler AJ. CD134/CD137 dual costimulation-elicited IFN-gamma maximizes effector T-cell
function but limits Treg expansion. Immunol Cell Biol. 2013;91(2):173-183.
doi pubmed - Razmkhah M, Abedi N, Hosseini A, Imani MT, Talei AR, Ghaderi
A. Induction of T regulatory subsets from naive CD4+ T cells after exposure to breast cancer
adipose derived stem cells. Iran J Immunol. 2015;12(1):1-15.
pubmed - Li R, Wang R, Zhong S, Asghar F, Li T, Zhu L, Zhu H.
TGF-beta1-overexpressing mesenchymal stem cells reciprocally regulate Th17/Treg cells by
regulating the expression of IFN-gamma. Open Life Sci. 2021;16(1):1193-1202.
doi pubmed - Wang K, Hu Z, Zhang C, Yang L, Feng L, Yang P, Yu H. SIRT5
contributes to colorectal cancer growth by regulating T cell activity. J Immunol Res.
2020;2020:3792409.
doi pubmed - Ghosh S, Roy K, Rajalingam R, Martin S, Pal C. Cytokines in
the generation and function of regulatory T cell subsets in leishmaniasis. Cytokine.
2021;147:155266.
doi pubmed - Bauer I, Grozio A, Lasiglie D, Basile G, Sturla L, Magnone
M, Sociali G, et al. The NAD+-dependent histone deacetylase SIRT6 promotes cytokine production
and migration in pancreatic cancer cells by regulating Ca2+ responses. J Biol Chem.
2012;287(49):40924-40937.
doi pubmed - Cai J, Wang D, Zhang G, Guo X. The role Of PD-1/PD-L1 axis
in treg development and function: implications for cancer immunotherapy. Onco Targets Ther.
2019;12:8437-8445.
doi pubmed - Pauken KE, Torchia JA, Chaudhri A, Sharpe AH, Freeman GJ.
Emerging concepts in PD-1 checkpoint biology. Semin Immunol. 2021;52:101480.
doi pubmed - Zhulai G, Oleinik E. Targeting regulatory T cells in
anti-PD-1/PD-L1 cancer immunotherapy. Scand J Immunol. 2022;95(3):e13129.
doi pubmed - Zhang Y, Liu Z, Tian M, Hu X, Wang L, Ji J, Liao A. The
altered PD-1/PD-L1 pathway delivers the 'one-two punch' effects to promote the Treg/Th17
imbalance in pre-eclampsia. Cell Mol Immunol. 2018;15(7):710-723.
doi pubmed - Karami Z, Mortezaee K, Majidpoor J. Dual
anti-PD-(L)1/TGF-beta inhibitors in cancer immunotherapy - Updated. Int Immunopharmacol.
2023;122:110648.
doi pubmed - Jiang L, Tang C, Gong Y, Liu Y, Rao J, Chen S, Qu W, et al.
PD-1/PD-L1 regulates Treg differentiation in pregnancy-induced hypertension.
Braz J Med Biol Res. 2018;51(8):e7334.
doi pubmed - Tahara M, Kondo Y, Yokosawa M, Tsuboi H, Takahashi S,
Shibayama S, Matsumoto I, et al. T-bet regulates differentiation of forkhead box protein 3+
regulatory T cells in programmed cell death-1-deficient mice. Clin Exp Immunol.
2015;179(2):197-209.
doi pubmed - Wang S, Li J, Xie J, Liu F, Duan Y, Wu Y, Huang S, et al.
Programmed death ligand 1 promotes lymph node metastasis and glucose metabolism in cervical
cancer by activating integrin beta4/SNAI1/SIRT3 signaling pathway. Oncogene.
2018;37(30):4164-4180.
doi pubmed - Song S, Tang H, Quan W, Shang A, Ling C. Estradiol initiates
the immune escape of non-small cell lung cancer cells via ERbeta/SIRT1/FOXO3a/PD-L1 axis. Int
Immunopharmacol. 2022;107:108629.
doi pubmed - Vaupel P, Mayer A. Hypoxia-driven adenosine accumulation: a
crucial microenvironmental factor promoting tumor progression. Adv Exp Med Biol.
2016;876:177-183.
doi pubmed - Whiteside TL. Regulatory T cell subsets in human cancer: are
they regulating for or against tumor progression? Cancer Immunol Immunother.
2014;63(1):67-72.
doi pubmed - Whiteside TL, Jackson EK. Adenosine and prostaglandin e2
production by human inducible regulatory T cells in health and disease. Front Immunol.
2013;4:212.
doi pubmed - Guo RS, Yu Y, Chen J, Chen YY, Shen N, Qiu M. Restoration of
brain acid soluble protein 1 inhibits proliferation and migration of thyroid cancer cells. Chin
Med J (Engl). 2016;129(12):1439-1446.
doi pubmed - Marsh LA, Carrera S, Shandilya J, Heesom KJ, Davidson AD,
Medler KF, Roberts SG. BASP1 interacts with oestrogen receptor alpha and modifies the tamoxifen
response. Cell Death Dis. 2017;8(5):e2771.
doi pubmed - Hartl M, Schneider R. A unique family of neuronal signaling
proteins implicated in oncogenesis and tumor suppression. Front Oncol. 2019;9:289.
doi pubmed - Zhou Q, Andersson R, Hu D, Bauden M, Kristl T, Sasor A,
Pawlowski K, et al. Quantitative proteomics identifies brain acid soluble protein 1 (BASP1) as a
prognostic biomarker candidate in pancreatic cancer tissue. EBioMedicine.
2019;43:282-294.
doi pubmed - Kropotova E, Klementiev B, Mosevitsky M. BASP1 and its N-end
fragments (BNEMFs) dynamics in rat brain during development. Neurochem Res.
2013;38(6):1278-1284.
doi pubmed - Hartl M, Puglisi K, Nist A, Raffeiner P, Bister K. The brain
acid-soluble protein 1 (BASP1) interferes with the oncogenic capacity of MYC and its binding to
calmodulin. Mol Oncol. 2020;14(3):625-644.
doi pubmed - Li L, Meng Q, Li G, Zhao L. BASP1 Suppresses cell growth and
metastasis through inhibiting Wnt/beta-Catenin pathway in gastric cancer. Biomed Res Int.
2020;2020:8628695.
doi pubmed - Ridder DA, Schindeldecker M, Weinmann A, Berndt K, Urbansky
L, Witzel HR, Heinrich S, et al. Key enzymes in pyrimidine synthesis, CAD and CPS1, predict
prognosis in hepatocellular carcinoma. Cancers (Basel). 2021;13(4):744.
doi pubmed - Zhang S, Hu Y, Wu Z, Zhou X, Wu T, Li P, Lian Q, et al.
Deficiency of carbamoyl phosphate synthetase 1 engenders radioresistance in hepatocellular
carcinoma via deubiquitinating c-Myc. Int J Radiat Oncol Biol Phys.
2023;115(5):1244-1256.
doi pubmed - Jiang MM, Zhao F, Lou TT. Assessment of significant pathway
signaling and prognostic value of GNG11 in ovarian serous cystadenocarcinoma.
Int J Gen Med. 2021;14:2329-2341.
doi pubmed - Li Q, Zhang Y, Jiang Q. MFAP5 suppression inhibits
migration/invasion, regulates cell cycle and induces apoptosis via promoting ROS production in
cervical cancer. Biochem Biophys Res Commun. 2018;507(1-4):51-58.
doi pubmed - Zhao L, Xu L, Hemmerich A, Ferguson NL, Guy CD, McCall SJ,
Cardona DM, et al. Reduced MFAP5 expression in stroma of gallbladder adenocarcinoma and its
potential diagnostic utility. Virchows Arch. 2021;478(3):427-434.
doi pubmed - Zhao L, Westerhoff M, Hornick JL, Krausz T, Antic T, Xiao
SY, Hart J. Loss of microfibril-associated protein 5 (MFAP5) expression in colon cancer stroma.
Virchows Arch. 2020;476(3):383-390.
doi pubmed - Aoki H, Yamamoto E, Takasawa A, Niinuma T, Yamano HO, Harada
T, Matsushita HO, et al. Epigenetic silencing of SMOC1 in traditional serrated adenoma and
colorectal cancer. Oncotarget. 2018;9(4):4707-4721.
doi pubmed - Gao H, Yan Z, Sun H, Chen Y. FoXA2 promotes esophageal
squamous cell carcinoma progression by ZEB2 activation. World J Surg Oncol.
2021;19(1):286.
doi pubmed - Han M, Li F, Zhang Y, Dai P, He J, Li Y, Zhu Y, et al. FOXA2
drives lineage plasticity and KIT pathway activation in neuroendocrine prostate cancer. Cancer
Cell. 2022;40(11):1306-1323.e1308.
doi pubmed - Zeng X, Yang X, Parhatsayim S, Abudoukelimu A, Shu Y, Zhao
Z. Forkhead box A2 promotes colorectal cancer progression and targets BCL2-associated X (BAX)
protein. Iran J Public Health. 2023;52(2):306-314.
doi pubmed - Oshita H, Nishino R, Takano A, Fujitomo T, Aragaki M, Kato
T, Akiyama H, et al. RASEF is a novel diagnostic biomarker and a therapeutic target for lung
cancer. Mol Cancer Res. 2013;11(8):937-951.
doi pubmed - Gurioli G, Martignano F, Salvi S, Costantini M, Gunelli R,
Casadio V. GSTP1 methylation in cancer: a liquid biopsy biomarker? Clin Chem Lab Med.
2018;56(5):702-717.
doi pubmed - Cui J, Li G, Yin J, Li L, Tan Y, Wei H, Liu B, et al. GSTP1
and cancer: Expression, methylation, polymorphisms and signaling (Review).
Int J Oncol. 2020;56(4):867-878.
doi pubmed - Liu J, Yu Z, Xiao Y, Meng Q, Wang Y, Chang W. Coordination
of FOXA2 and SIRT6 suppresses the hepatocellular carcinoma progression through ZEB2 inhibition.
Cancer Manag Res. 2018;10:391-402.
doi pubmed - Wang T, Liu X, Wang T, Zhan L, Zhang M. BASP1 expression is
associated with poor prognosis and is correlated with immune infiltration in gastric cancer.
FEBS Open Bio. 2023;13(8):1507-1521.
doi pubmed - Pan X, Xu X, Wang L, Zhang S, Chen Y, Yang R, Chen X, et al.
BASP1 is a prognostic biomarker associated with immunotherapeutic response in head and neck
squamous cell carcinoma. Front Oncol. 2023;13:1021262.
doi pubmed - Wang J, Xia S, Zhao J, Gong C, Xi Q, Sun W. Prognostic
potential of secreted modular calcium-binding protein 1 in low-grade glioma. Front Mol Biosci.
2021;8:666623.
doi pubmed - Heyliger SO, Soliman KFA, Saulsbury MD, Reams RR. Prognostic
Relevance of ZNF844 and Chr 19p13.2 KRAB-zinc finger proteins in clear cell renal carcinoma.
Cancer Genomics Proteomics. 2022;19(3):305-327.
doi pubmed - Luo L, Wang Y, Feng Q, Zhang H, Xue B, Shen J, Ye Y, et al.
Recombinant protein glutathione S-transferases P1 attenuates inflammation in mice. Mol Immunol.
2009;46(5):848-857.
doi pubmed - Lebrec H, Ponce R, Preston BD, Iles J, Born TL, Hooper M.
Tumor necrosis factor, tumor necrosis factor inhibition, and cancer risk. Curr Med Res Opin.
2015;31(3):557-574.
doi pubmed - Gaul DS, Calatayud N, Pahla J, Bonetti NR, Wang YJ, Weber J,
Ambrosini S, et al. Endothelial SIRT6 deficiency promotes arterial thrombosis in mice.
J Mol Cell Cardiol. 2023;174:56-62.
doi pubmed - Hong OY, Jang HY, Lee YR, Jung SH, Youn HJ, Kim JS.
Inhibition of cell invasion and migration by targeting matrix metalloproteinase-9 expression via
sirtuin 6 silencing in human breast cancer cells. Sci Rep. 2022;12(1):12125.
doi pubmed - Gloushankova NA, Zhitnyak IY, Rubtsova SN. Role of
epithelial-mesenchymal transition in tumor progression. Biochemistry (Mosc).
2018;83(12):1469-1476.
doi pubmed
This
article is distributed under the terms of the Creative Commons Attribution Non-Commercial 4.0
International License, which permits unrestricted non-commercial use, distribution, and
reproduction in any medium, provided the original work is properly cited.
World
Journal of Oncology is published by Elmer Press Inc.