| World Journal of Oncology, ISSN 1920-4531 print, 1920-454X online, Open Access |
| Article copyright, the authors; Journal compilation copyright, World J Oncol and Elmer Press Inc |
| Journal website https://wjon.elmerpub.com |
Original Article
Volume 16, Number 2, April 2025, pages 161-172
Actionable Mutations and Survival Rates in Non-Small Cell Lung Cancer
Talal S. Al-Qaisia, Berjas Abumsimira, d, Maher Sughayerb, Yassine Kasmic
aDepartment of Medical Laboratory Sciences, Pharmacological and Diagnostic Research
Centre (PDRC), Faculty of Allied Medical Sciences, Al-Ahliyya Amman University (AAU), Amman
19328, Jordan
bDepartment of Pathology, King Hussein Cancer Center, Amman,
Jordan
cJohann Heinrich von Thunen Institute, Braunschweig 38116,
Germany
dCorresponding Author: Berjas Abumsimir, Department of Medical
Laboratory Sciences, Pharmacological and Diagnostic Research Centre (PDRC), Faculty of Allied
Medical Sciences, Al-Ahliyya Amman University (AAU), Amman 19328, Jordan
Manuscript submitted January 3, 2025, accepted February 19, 2025, published online March 7,
2025
Short title: Actionable Mutations and Survival Rates of NSCLC
doi:
https://doi.org/10.14740/wjon2531
| Abstract | ▴Top |
Background: In Jordan, lung cancer ranks as the second most common tumor, and there is an urgent need to explore the genetic landscape of lung cancer. This study aimed to identify the actionable mutations in lung cancer samples in Jordanians by targeted next-generation sequencing (NGS) and to investigate the correlations with clinical and pathological parameters.
Methods: Totally, 121samples were prepared for NGS by DNA extractions from formalin-fixed paraffin-embedded (FFPE) blocks, followed by library preparation using the AmpliSeq Colon and Lung panel, which covers mutational hot spot regions for 22 cancer genes.
Results: Amongst 121 patients, 88% of those treated for non-small lung carcinoma were successfully analyzed; 35 (29%) carried one mutation or more in actionable genes (KRAS, EGFR, ALK, BRAF, and MET). There are no significant differences between actionable mutation carriers and non-carriers concerning histological tumor type, tumor stage, metastasis, smoking habits, and gender. However, the analysis of survival probabilities revealed lower survival times for females compared to males, as well as for those patients who had metastasis events, smoking, or relapse after treatment.
Conclusions: The type and rates of mutations detected for lung tumors in Jordan are relatively similar to those found in other populations previously studied, although some differences exist. However, lung tumors in Jordan require new customized treatment prescriptions based on prior genetic studies, as part of the hoped-for trend toward precision medicine.
Keywords: Lung cancer; Actionable mutations; Next-generation sequencing; Non-small cell lung cancer; Jordan
| Introduction | ▴Top |
Lung cancer remains a significant public health concern, and efforts to prevent and manage it are crucial. According to the World Cancer Research Fund, the latest new data of lung cancer worldwide for 2022, there were 2,480,675 new cases of lung cancer globally. USA and China lead the top 10 countries with the highest rates of lung cancer incidence: China with 1,060,584 cases and USA with 226,033 cases. There were 1,817,469 deaths from lung cancer globally, with 733,291 deaths in China and 127,653 deaths in the USA [1].
The age-standardized rate (ASR) of lung cancer in the Middle East and North African (MENA) region is generally lower compared to international rates. The ASR varies across countries, ranging from 4.2 cases per 100,000 in Yemen to 23 cases per 100,000 in Lebanon. Lung cancer ranks among the top 10 most common cancers in most MENA countries. In Jordan, with a population of approximately 10 million, there were 10,898 new cancers cases reported. Lung cancer accounted for 1,141 cases (10.5% of all cancer cases) and ranked as the second most common cancer type, with 1,022 deaths (5.6%), and 1,049 for 5-year prevalent cases [2].
Lung cancer mutations arise from changes in specific genes, and these mutations play a key role in the development and progression of the tumor. The mutations could be inherited mutations (germline), which are passed from parents to their children. However, they account for only 5-10% of lung cancer cases. Acquired mutations (somatic), occur during a person’s lifetime due to exposure to various factors such as tobacco smoke. Smoking is the leading cause of lung cancer, responsible for approximately 90% of cases. Other factors include air pollution, chemicals, radon gas, and ultraviolet (UV) radiation [3].
The most common lung cancer mutations are EGFR (epidermal growth factor receptor) mutations, which are more prevalent in certain types of lung cancer, especially in non-smokers and people of Asian descent. Mutations in TP53 occur in about 43% of lung squamous cell carcinomas (SCCs) and 35% of lung adenocarcinomas. KRAS mutations were found in approximately 30% of non-small cell lung cancers (NSCLCs), KRAS mutations are more common in smokers. ALK and ROS1 mutations, along with EGFR and other genes, are relevant for targeted therapies in specific cases [4, 5].
These days, lung cancer treatment aims to manage late-stage cancer, and many drugs are sensitive to specific mutations in lung cancer. Precision medicines aim to treat patients who suffer from cancer and have special mutations in genes such as EGFR, KRAS, ALK, and other genes [6].
Studies that have focused on lung cancer mutations in Jordanian population are few, and the medical sector in Jordan is interested in moving to precision medicine methods in treating cancer, including lung cancer. Therefore, it is important to conduct more studies that examine genetic mutations in lung cancer patients in Jordan in a specialized and continuous manner, and to compare them with populations that belong to the same genetic pool.
This study aims to fill a gap in this field and is an attempt to shed light on the general genetic landscape of lung cancer in Jordan, contributing to the fight against lung cancer in cooperation with the King Hussein Cancer Center (KHCC), the leading center in Jordan in this field. This study aims primarily to identify actionable mutations in lung cancer samples from Jordanians and, secondly, to investigate the correlation between genetic mutations and clinical and pathological parameters.
| Materials and Methods | ▴Top |
Population and samples
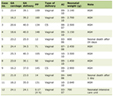
Formalin-fixed paraffin-embedded (FFPE) samples from 140 lung cancer patients at the Pathology Department of KHCC, Jordan in 2021 were eligible for mutational analysis, and 121 samples were successfully analyzed. The Pathology Department at KHCC was responsible for obtaining the samples, arrangement, and histological classifications. All clinical and pathological data were recorded. Histological assessment and pathological staging were performed according to the World Health Organization (WHO) classification, and American Joint Committee on Cancer (AJCC) guidelines, respectively [7, 8]. The ethical approval was obtained in accordance with the regulation of the Institutional Review Board (IRB) at KHCC (No. KHCC 81). All patients consented to participate in the study, and their information was handled anonymously according to ethical standards. Among 121 patients, 83% were males, and 73% were above 60 years or older. Eighty-eight percent of the patients were diagnosed with NSCLC, and three patients were diagnosed with small cell lung cancer (SCLC). Fifty-six percent of all patients were at the T1 or T2 clinical stage of tumor. The vast majority (73%) of them had primary lung cancer, and 7% had metastasis events. Seventy-two percent of all patients are smokers, and 99% of them have been treated for NSCLC (Table 1).
 Click to view |
Table 1. Clinical and Pathological
Characteristics of Lung Cancer Patients |
DNA extraction
DNA was extracted by the Zymo DNA Mini Kit (Zymo, USA) according to the manufacturer’s protocol. DNA quantification was performed using a fluorometric method, such as Qubit or PicoGreen. If enough DNA is available, the DNA samples were diluted to a concentration of about 20 - 50 ng/µL using low Tris-EDTA (TE). Then the diluted DNA samples were re-quantified using the same fluorometric quantification method to the desired final concentration using low TE.
Next-generation sequencing (NGS)
Sequencing libraries were prepared from 10 ng of DNA samples using the Ion AmpliSeq Colon and Lung Cancer Panel V2®, together with the Ion AmpliSeq Library Preparation Kit (Thermo Fisher Scientific), according to the manufacturer’s instructions. The colon and lung cancer panel contains 504 mutational hot spots (overall 14.6 kb) of 22 genes: AKT1, ALK, BRAF, CTNNB1, DDR2, EGFR, ERBB2, ERBB4, FBXW7, FGFR1, FGFR2, FGFR3, KRAS, MAP2K1, MET, NOTCH1, NRAS, PIK3CA, PTEN, SMAD4, STK11, and TP53. The libraries were quantified with the Qubit 2.0 Fluorometer, using the Qubit dsDNA HS Assay Kit. Templates were prepared and enriched using the Ion OneTouch 2 system and the Ion PGM Template OT2 200 Kit. Library quality control was accomplished by the Ion Sphere Quality Control Kit according to the manufacturer’s instructions, confirming that 10-30% of template-positive ion sphere particles (ISPs) were targeted in the emPCR reaction. Sequencing was performed with the Ion PGM Sequencing 200 Kit v2, the Ion 316v2 chip, and the Ion Torrent PGM instrument (Ion PGM™, Thermo Fisher Scientific) with the Ion PGM™ Hi-Q View™ Sequencing Kit (Thermo Fisher Scientific).
Variant annotation and data analysis
The raw sequencing data were aligned to the hg39 and analyzed with the Basepair tech server after eliminating low-quality reads and trimming adapter sequences. Mutations were detected using variant caller tools in the Basepair tech server from the reference sequence using default settings, i.e., a quality score of minimum 6, a relative read quality of minimum 6.5, coverage of minimum 6 for SNP/COSMIC variant and 15 for insertion-deletion mutation (indel), and strand bias maximum of 95% for SNP/COSMIC variant and 90% for indel. To find pathogenic variations, mutations that did not affect the protein coding regions (intronic, untranslated region (UTR) variations) were excluded. All detected variants were manually reviewed with the Mega X software. This analysis was enriched by comparison between variant caller outputs with ClinVar and dSNP-NCBI databases. Further data filtering was conducted to include only those variants that had a frequency of variant reads with a minimum of 1% or less. The coverage analysis was performed using the coverage analysis plugin (v4.0) (Thermo Fisher Scientific). Statistical analyses were performed with Minitab version 18 (Coventry, UK). Fisher’s exact test was used to compare mutation-positive and negative groups with categorical features. An independent t-test was used for the comparison of mutation statuses and age. The two-sided P values < 0.05 were considered statistically significant. The survival rates were generated using the Kaplan-Mier curve method by the SRplot server [9]. The survival rates were compared according to gender, metastasis event, relapse during treatment, and whether the lung tumor was primary or secondary.
| Results | ▴Top |
The panel AmpliSeq Cancer HotSpot Panel v2 for Illumina was used to target the hot spot of 22 genes mentioned in the methodology section. All patients have at least one mutation in one gene or more from the target panel. For a more comprehensive analysis, the genes with synonymous and nonsynonymous mutations were considered, and the synonymous mutation only.
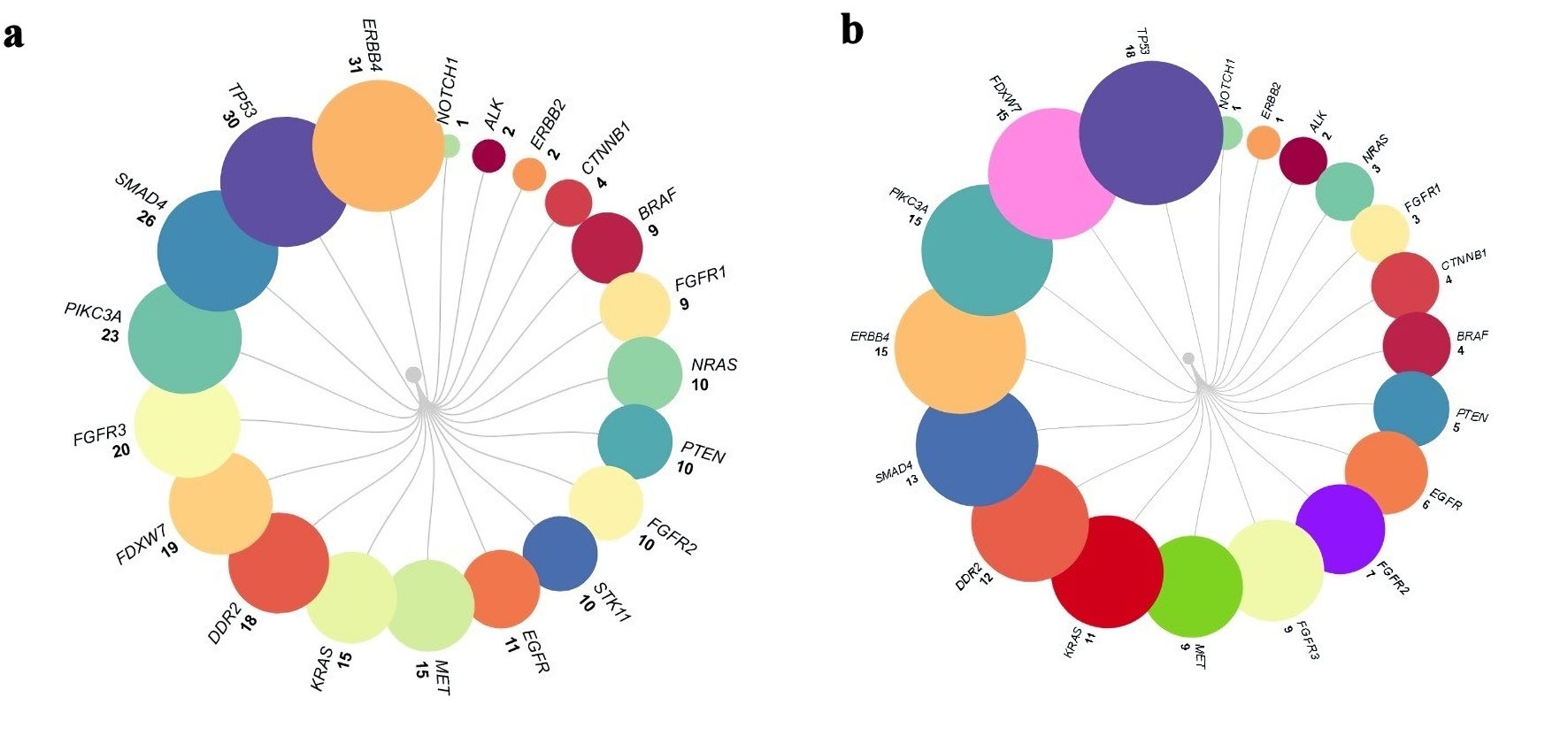
The most frequent mutations were estimated as the first step of analysis. The genes with 10 variants or more were: NRAS, PTEN, FGFR1, STK11, EGFR, MET, KRAS, DDR2, FDXW7, FGFR3, PIKC3A, SMAD4, TB53, and ERBB4, with 10, 10, 10, 11, 15, 18, 19, 20, 23, 26, 30 and 31 variants, respectively. While when the synonymous variants were excluded, the genes with 10 variants or more were as follows: KRAS, DDR2, SMAD4, ERBB4, PIKC3A, FDXW7, and TP53, with 11, 12, 13, 15, 15 and 18 nonsynonymous variants, respectively. After this initial analysis, we went deep to explore specific variants for each gene, focusing on the actionable (druggable) genes. The mutations detected for each gene are presented, including synonymous variants (Fig. 1a), and excluding synonymous variants arranged in ascending order (Fig. 1b).
 Click for large image |
Figure 1. The mutations for each gene including synonymous variants (a), and excluding synonymous variants arranged in ascending order (b). |
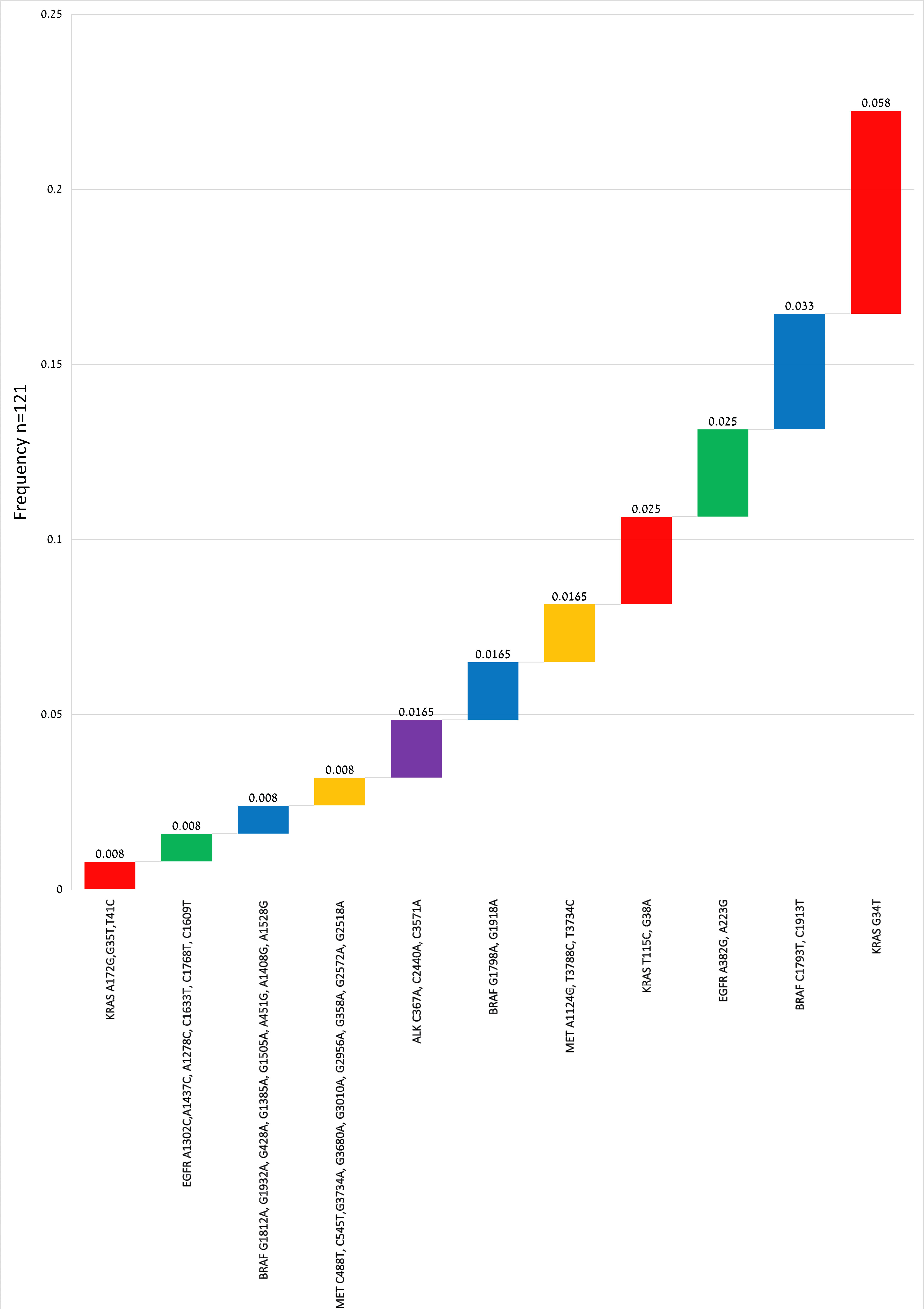
The most frequent variant in KRAS was c.G34T:p.G12C found in seven patients (0.058), followed by c.G38A:p.G13D and c.T115C:p.S39P in three patients, with frequency of 0.025. In EGFR, c.A382G:p.N128D, and c.A223G:p.N75D were the most frequent variants (0.025) found in three patients. In ALK, c.C367A:p.P123T, c.C2440A:p.P814T, and c.C3571A:p.P1191T were detected in two patients (0.0165). In BRAF, c.C1793T:p.A598V, and c.C1913T:p.A638V were observed in four patients (0.033), and c.G1798A:p.V600M in two patients. The other BRAF mutations were observed in one patient for each variant. Finally MET variants c.A1124G:p.N375S, and c.T3788C:p.L1263P, were observed in two samples. The most actionable gene variants excluding synonymous mutations and subsequent frequencies, were given in Figure 2.
 Click for large image |
Figure 2. Mutations of actionable genes detected in 121 lung cancer patients (color-coded by gene). Reference genome database: GRCh38, and Human Genome Variation Society (HGVS) nomenclature. |
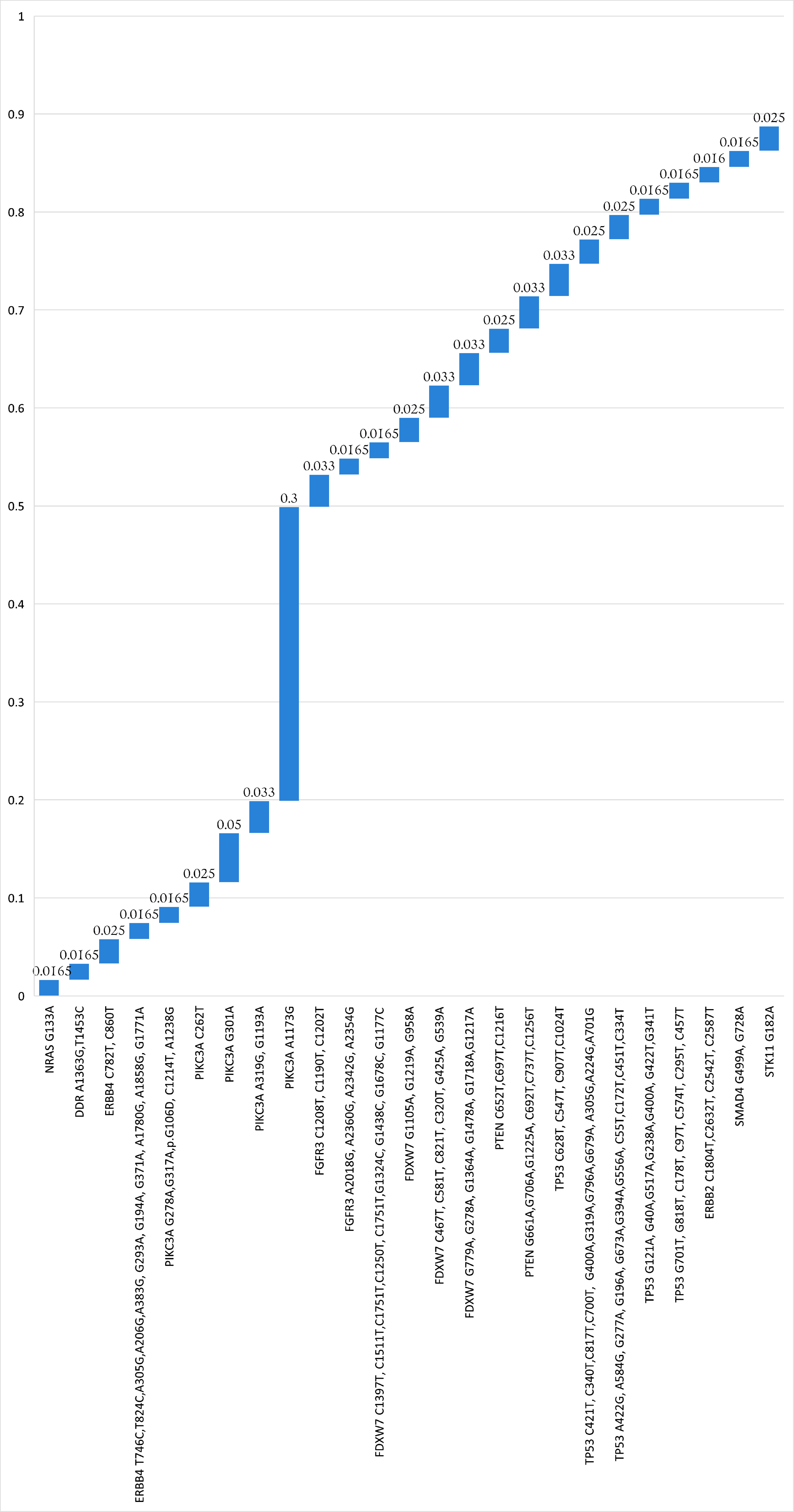
For comprehensive elucidation of the genetic landscape of lung cancer, other gene mutations were analyzed excluding synonymous ones. The variants observed in two patients, or more, are presented in Figure 3, and the variants detected in one patient only were included here (Supplementary Material 1, wjon.elmerpub.com). Most variants were repeated in two or three patients (frequency rates: 0.0165 and 0.025, respectively). A notable missense mutation in PIKC3A exon7: c.A1173G:p.I391M was found in 37 patients (0.30), which seems very high in this population and needs further validation. Other missense mutations in the same gene but in exon2: c.G301A:p.V101I, and in exon7: c.G1193A:p.R398H, were found in six patients (0.05), and four patients (0.033), respectively. In FGFR3 exactly in exon9, c.C1208T:p.P403L, c.C1190T:p.P397L and c.C1202T:p.P401L were found in four patients (0.033). Four mutations in four patients each at the same exon10 of FDXW7 were observed, namely: c.G1364A:p.C455Y, c.G1478A:p.C493Y, c.G1718A:p.C573Y and c.G1217A:p.C406Y. PTEN has more higher mutational events in exons 6, 7, and 8 as follows: exon6: c.G661A:p.D221N, c.C692T:p.P231L, exon7:c.G706A:p.D236N, c.C737T:p.P246L, and exon8: c.G1225A:p.D409N, c.C1256T:p.P419L.
 Click for large image |
Figure 3. Mutations of other actionable genes detected in two patients or more. Reference genome database: (GRCh38), and Human Genome Variation Society (HGVS) nomenclature. |
We went further to analyze indels. Indels are influential in clinical NGS, as they are associated with the steering mechanism underlying many tumors. High-frequent frameshift insertion were detected in NOTCH in 49 patients (0.40), namely: c.2331_2332insTG: p. L778Cfs*2, c.4620_4621insTG:p.L1541Cfs*2, 4734_4735insTG:p.L1579Cfs*2, and 4614_4615insTG:p.L1539Cfs*2. Frameshift insertions c.1720dupG: p. A574Gfs*10 and c.1798dupG: p. A600Gfs*10 were detected in seven patients (0.058). For NRAS, a non-frameshift insertion c.26_27insGGT: p.V9_G10insV was observed in four patients (0.033). The rest of the indels were identified in one, two, or three patients at most, as detailed in Table 2.
 Click to view |
Table 2. Indels Variations Detected in All
Genes Hot Spots of Panel Analyzed |
Amongst 121 patients successfully analyzed, 35 (29%) carried one mutation or more in actionable genes (KRAS, EGFR, ALK, BRAF, and MET). For every tumor characteristic, the number and percentages of patients who are carrying mutations and non-carriers were recorded. Among actionable mutations carriers, 28 (80%) were diagnosed with NSCLC subtypes: 23 (66%) patients have lung adenocarcinoma, and five (14%) have squamous cell carcinoma. Notably, none of them were diagnosed with SCLC or large-cell neuroendocrine carcinoma, with no significant differences between carriers and non-carriers (P value = 0.574) according to tumor histological type.
Regarding the pathological tumor stage, 13 (37%) of the mutation, carriers were at T3 or T4 of the tumor stage, in a close manner, 26 (30%) of the non-carriers were at the same stages (P value = 0.438). Twenty-one of those were diagnosed at 60 years or less, carrying one or more actionable mutations, while 58 non-carriers were more than 60 years old at the time of diagnosis (P = 0.946). Among carriers of these mutations found, 24 (69%) have no tumor metastasis events, and 24 (69%) have primary tumors; while among non-carriers, the ratios were 60 (70%) and 69 (80%), respectively, with no significant differences for both parameters. Among actionable mutations, 21 carriers were smokers, and 51 non-carriers were smokers, with no significant differences (P = 0.484). Tumor characteristics of lung cancer patients with actionable mutations, both carriers and non-carriers are shown in Table 3.
 Click to view |
Table 3. Correlation of Clinical and
Pathological Parameters Between Carriers and Non-Carriers of Actionable
Variants |
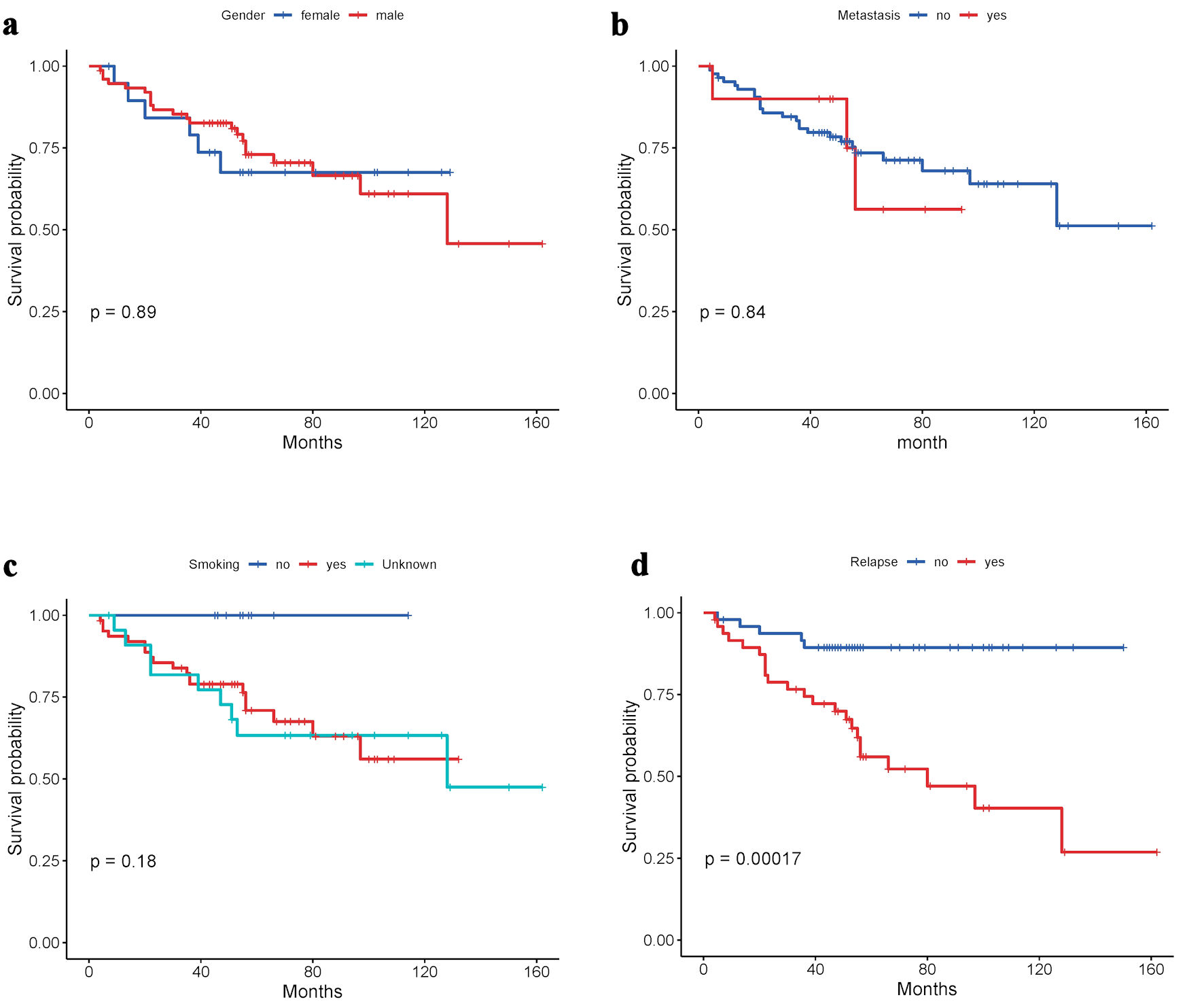
Kaplan Meier curves were generated to estimate survival patterns concerning clinical and pathological parameters, namely: gender, metastasis events, smoking, and relapse (Fig. 4). The analyses revealed lower survival times for females compared to males, with survival probability decreasing for female lung cancer patients after 80 months (P = 0.89). Lower survival rates were observed in those patients who had metastasis events or relapses after treatment in comparison with patients without metastasis or relapse events, with survival probability decreasing very early, starting from 40 - 60 months (P = 0.84 and P = 0.00017, respectively). The survival probability analysis for smoking habits showed decreasing survival time probability for smokers in comparison with non-smokers from early months up to maximum 160 months.
 Click for large image |
Figure 4. Kaplan-Meier survival curves of patients according to (a) gender, (b) metastasis events, (c) smoking, and (d) relapse. |
| Discussion | ▴Top |
The Jordanian population is considered a typical Arabian population, and therefore an example of the genetic pool in this region of the world. This genetic influence is reflected as a risk factor for many diseases, including cancer, and in the case of our study, lung cancer. The world entered the era of precision medicine some time ago, and this field is developing and becoming more important for the precise treatment of lung tumors based on the genetic profile of each patient. This means specialized genetic studies are needed, which shed light on the genetic landscape of any population. In our study here, we aimed to explore the genetic landscape of lung cancer in Jordan as the first step toward applying precision medicine for tumor treatment in Jordan. For a long time, Jordan has needed such a study targeting specific mutations in specific genes that are universally recommended for lung cancer. This is the first study in Jordan focused specifically on the genetic portrait of lung tumors in the Jordanian population.
In our study, KRAS G12C frequency was estimated at around 6%, in comparison with a frequency of 8.9-19.5% in the USA, 9.3-18.4% in Europe, 6.9-9.0% in Latin America, and 1.4-4.3% in Asia [10]. The frequency of KRAS G12C in the Jordanian population is lower than in the USA, higher than in Asian, and semi-equal Latin America ancestry and some European populations. To date, there are no applicable strategies targeting KRAS mutations, except for Sotorasib, which is an inhibitor of the RAS GTPase family and has recently been approved to target the KRAS (G12C). KRAS G13 mutation frequencies, as recorded previously, are less than 1% [11]. Our findings indicate this frequency to be approximately 2.5% in Jordanian non-small-cell lung cancer patients. The reported EGFR mutation frequencies in patients with lung adenocarcinoma differ broadly between different populations globally, ranging from 13.5% of patients with NSCLC and 14.7% of patients with lung adenocarcinomas in European countries, and about 50% in Asian populations [12, 13]. In our samples, no notable mutation with recommended therapy was found. The most frequent missense variants were N128D and N75D (2.5%), which were not reported before. ALK gene fusion and rearrangements are more commonly associated with therapy response. In contrast, ALK missense variants are still not targetable in lung tumors. In our study, the most common missense mutations were P123T, P814T, and P1191T (frequency 1.65%). BRAF V600E, reported in some populations [14], was not detectable in our study, although the mutation V600M was detected in two patients (1.65%). Furthermore, the A598V was detected in 3.3% of lung cancer patients in Jordan. MET N375S mutation was reported before in lung cancer patients somewhere [15]. Indeed, this mutation was observed in our study of NSCLC patients with a frequency of 1.65%. ERBB2 (erythroblastic oncogene B) is mutated in approximately 1-4% of lung adenocarcinoma cases [16]. In this study, the highest ERBB2 frequency was 2.5%, which is close to that range. A notable high-frequency (30%) mutation in exon7: c. A1173G:p.I391M of PIKC3A, needs more investigations, especially since this mutation has been reported in many types of human tumors other than lung adenocarcinoma. Clinical research recognized that lung tumors having mutated PTEN are linked with low survival rates. PTEN has been investigated as a promising target for anti-cancer therapy [17]. Despite previous reports revealing that ALK (anaplastic lymphoma kinase) gene rearrangement occurs in 3-6% of all NSCLC [18], our finding reported less than 1% of ALK frameshift insertion and less than 2% of point mutations. Our finding also confirmed the role of NOTCH frameshift mutations in lung cancer development [19], since about 40% of lung cancer patients have one or more frameshift insertion mutations (Table 2).
In our group, around 80-85% of lung cancer cases are NSCLC; the rest are usually diagnosed as SCLC. In our sample, the ratio of NSCLC was more than 87%. We did not find any significant correlation between actionable mutations carriers (n = 35), non-carriers (n = 86), and clinical and pathological characteristics regarding cancer histological types, gender, tumor stage, metastasis, lung tissues-initiation of the tumor, or smoking habits. This is due to the random nature of the mutations, which may be inherited or de novo mutations, as there are no studies that limit the occurrence of these mutations to a specific histological type of lung tumor, or in males but not females. However, there is accumulated evidence that smoking could damage lung cells’ DNA, along with other environmental factors such as UV radiation and radioactive substances [20]. However, the sample size and the lack of information may have affected the significant differences between these two groups of lung cancer patients.
To explore the survival rate of our lung cancer patients, the data were analyzed according to metastasis events, gender, relapse, or primary tumor type. These four factors effectively influence survival rates. Smoking habits affected the probability of survival times by two-fold, starting from the early month after treatment up to 160 months of survival. The physiological differences between males and females, as observed from the original data, affected the clinical relapse in females more than males (P = 0.89). Similarly, metastasis events of tumors contributed to aggressivity and shorter life span after diagnosis, regardless of gender (P = 0.84). The initiation of a tumor in the lung (primary) or metastasis into the lung (secondary) is another critical factor for survival, as it makes the life span after diagnosis shorter when the tumors were secondary, despite no significant differences (P = 0.5). Finally, the relapse of some patients significantly impacted survival rate in this subset of lung cancer patients (P = 0.00017) (Fig. 4).
Conclusions
It is self-evident that the genetic landscape of lung tumor mutations in Jordan is close to that in Arab countries, especially nearby ones, as it is considered a typical Arabian population. At the same time, and to a lesser extent, the genetic landscape of lung tumors in Jordan is not too far from the genetic landscape of lung tumors in populations previously explored, especially in the Mediterranean and Europe. However, it constitutes a unique case of the genetic pool that deserves the design of specialized therapeutic plans, which suit the nature of the genetic mutations in it. Following this study, other confirmatory studies are needed to validate the genetic landscape of lung tumors in Jordan. It is time for lung cancer treatments in Jordan to going deep in the era of precision medicine, which will make a big difference in improving the lives of patients and increasing the survival rate of these patients. This places a double effort on the treatment foundations and the use of specific genetic investigations during diagnosis and treatment.
New genetic technologies, such as NGS and subsequent analysis and variant calling, have limitations in detecting various variants. Therefore, studies focusing specific genes such as KRAS or EGFR alterations, are needed in Jordanian population. It is possible to expand the number of samples beyond what is currently available to us, provide more comprehensive clinical and demographic information, and conduct similar studies to reveal the environmental and inherited risk factors that lead to the emergence and development of lung tumors in Jordan.
| Supplementary Material | ▴Top |
Suppl 1. Mutations in other genes that are observed in one sample only.
Acknowledgments
The authors would like to thank the Deanship of Scientific Research at Al-Ahliyya Amman University for its continuous support and follow-up from the beginning of the study until the completion of writing this study. The researchers would also like to thank the KHCC for their positive cooperation during the study period, represented by the presidency and all workers of the hospital and center.
Financial Disclosure
The study was funded by the Deanship of Scientific Research at Al-Ahliyya Amman University (No. 2022-2021/30/2).
Conflict of Interest
None to declare.
Informed Consent
Not applicable.
Author Contributions
BA and TA conceived and designed the study. BA, TA, and MS optimized the experimental approach. BA, TA and MS performed the experiments. BA, TA, YK and MS analyzed all the sequencing data and performed the statistical analyses. TA and MS managed sample collection, processed the data, and supervised the storage of all samples. Proofreading and technical assistance were carried out by BA, TA, MS and YK, who wrote the manuscript. All authors approved the final manuscript submission.
Data Availability
All data required are included in this manuscript. Raw data can be requested from authors directly.
Abbreviations
SCC: squamous cell carcinomas; NSCLC: non-small cell lung cancers; ASR: age-standardized rate; MENA: Middle East and North African; SNP: single nucleotide polymorphisms
| References | ▴Top |
- https://www.wcrf.org/cancer-trends/lung-cancer-statistics/ (retrieved July 2024).
- Jazieh AR, Algwaiz G, Errihani H, Elghissassi I,
Mula-Hussain L, Bawazir AA, Gaafar R. Lung cancer in the Middle East and North Africa Region.
J Thorac Oncol. 2019;14(11):1884-1891.
doi pubmed - Wadowska K, Bil-Lula I, Trembecki L, Sliwinska-Mosson M.
Genetic markers in lung cancer diagnosis: a review. Int J Mol Sci.
2020;21(13):4569.
doi pubmed - Tumbrink HL, Heimsoeth A, Sos ML. The next tier of EGFR
resistance mutations in lung cancer. Oncogene. 2021;40(1):1-11.
doi pubmed - Ghimessy A, Radeczky P, Laszlo V, Hegedus B, Renyi-Vamos F,
Fillinger J, Klepetko W, et al. Current therapy of KRAS-mutant lung cancer. Cancer Metastasis
Rev. 2020;39(4):1159-1177.
doi pubmed - Lamberti G, Andrini E, Sisi M, Rizzo A, Parisi C, Di
Federico A, Gelsomino F, et al. Beyond EGFR, ALK and ROS1: Current evidence and future
perspectives on newly targetable oncogenic drivers in lung adenocarcinoma. Crit Rev Oncol
Hematol. 2020;156:103119.
doi pubmed - Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM,
Beasley MB, Chirieac LR, et al. The 2015 World Health Organization Classification of lung
tumors: impact of genetic, clinical and radiologic advances since the 2004 classification.
J Thorac Oncol. 2015;10(9):1243-1260.
doi pubmed - Travis WD, Committee IS. Reporting lung cancer pathology
specimens. Impact of the anticipated 7th Edition TNM classification based on recommendations of
the IASLC Staging Committee. Histopathology. 2009;54(1):3-11.
doi pubmed - Tang D, Chen M, Huang X, Zhang G, Zeng L, Zhang G, Wu S, et
al. SRplot: A free online platform for data visualization and graphing. PLoS One.
2023;18(11):e0294236.
doi pubmed - Lim TKH, Skoulidis F, Kerr KM, Ahn MJ, Kapp JR, Soares FA,
Yatabe Y. KRAS G12C in advanced NSCLC: Prevalence, co-mutations, and testing. Lung Cancer.
2023;184:107293.
doi pubmed - Huang L, Guo Z, Wang F, Fu L. KRAS mutation: from
undruggable to druggable in cancer. Signal Transduct Target Ther. 2021;6(1):386.
doi pubmed - Grosse A, Grosse C, Rechsteiner M, Soltermann A. Analysis of
the frequency of oncogenic driver mutations and correlation with clinicopathological
characteristics in patients with lung adenocarcinoma from Northeastern Switzerland. Diagn
Pathol. 2019;14(1):18.
doi pubmed - Ramlau R, Cufer T, Berzinec P, Dziadziuszko R, Olszewski W,
Popper H, Bajcic P, et al. Epidermal growth factor receptor mutation-positive non-small-cell
lung cancer in the real-world setting in central Europe: the INSIGHT study. J Thorac
Oncol. 2015;10(9):1370-1374.
doi pubmed - Toth LJ, Mokanszki A, Mehes G. The rapidly changing field of
predictive biomarkers of non-small cell lung cancer. Pathol Oncol Res. 2024;30:1611733.
doi pubmed - Guerin C, Tulasne D. Recording and classifying MET receptor
mutations in cancers. Elife. 2024;13:e92762.
doi pubmed - Sharma M, Dewan A, Diwan H, Nathany S, Batra U. A narrative review of ERBB2 in non-small cell lung carcinoma. Cancer Research, Statistics, and Treatment. 2022;5(1):97-104.
- Alvarez-Garcia V, Tawil Y, Wise HM, Leslie NR. Mechanisms of
PTEN loss in cancer: It's all about diversity. Semin Cancer Biol. 2019;59:66-79.
doi pubmed - Inamura K, Takeuchi K, Togashi Y, Nomura K, Ninomiya H, Okui
M, Satoh Y, et al. EML4-ALK fusion is linked to histological characteristics in a subset of lung
cancers. J Thorac Oncol. 2008;3(1):13-17.
doi pubmed - Galluzzo P, Bocchetta M. Notch signaling in lung cancer.
Expert Rev Anticancer Ther. 2011;11(4):533-540.
doi pubmed - Gibbons DL, Byers LA, Kurie JM. Smoking, p53 mutation, and
lung cancer. Mol Cancer Res. 2014;12(1):3-13.
doi pubmed
This
article is distributed under the terms of the Creative Commons Attribution Non-Commercial 4.0
International License, which permits unrestricted non-commercial use, distribution, and
reproduction in any medium, provided the original work is properly cited.
World
Journal of Oncology is published by Elmer Press Inc.