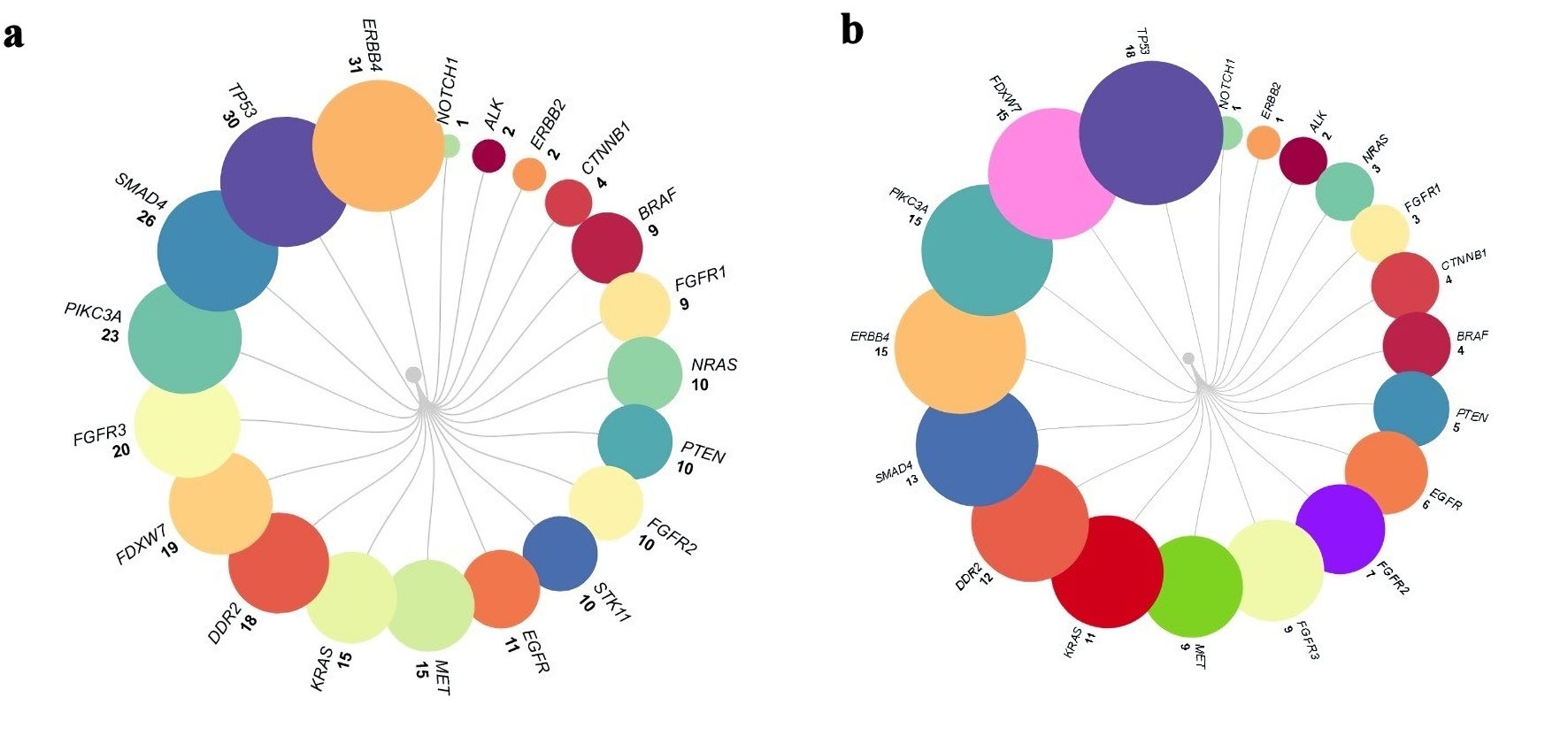
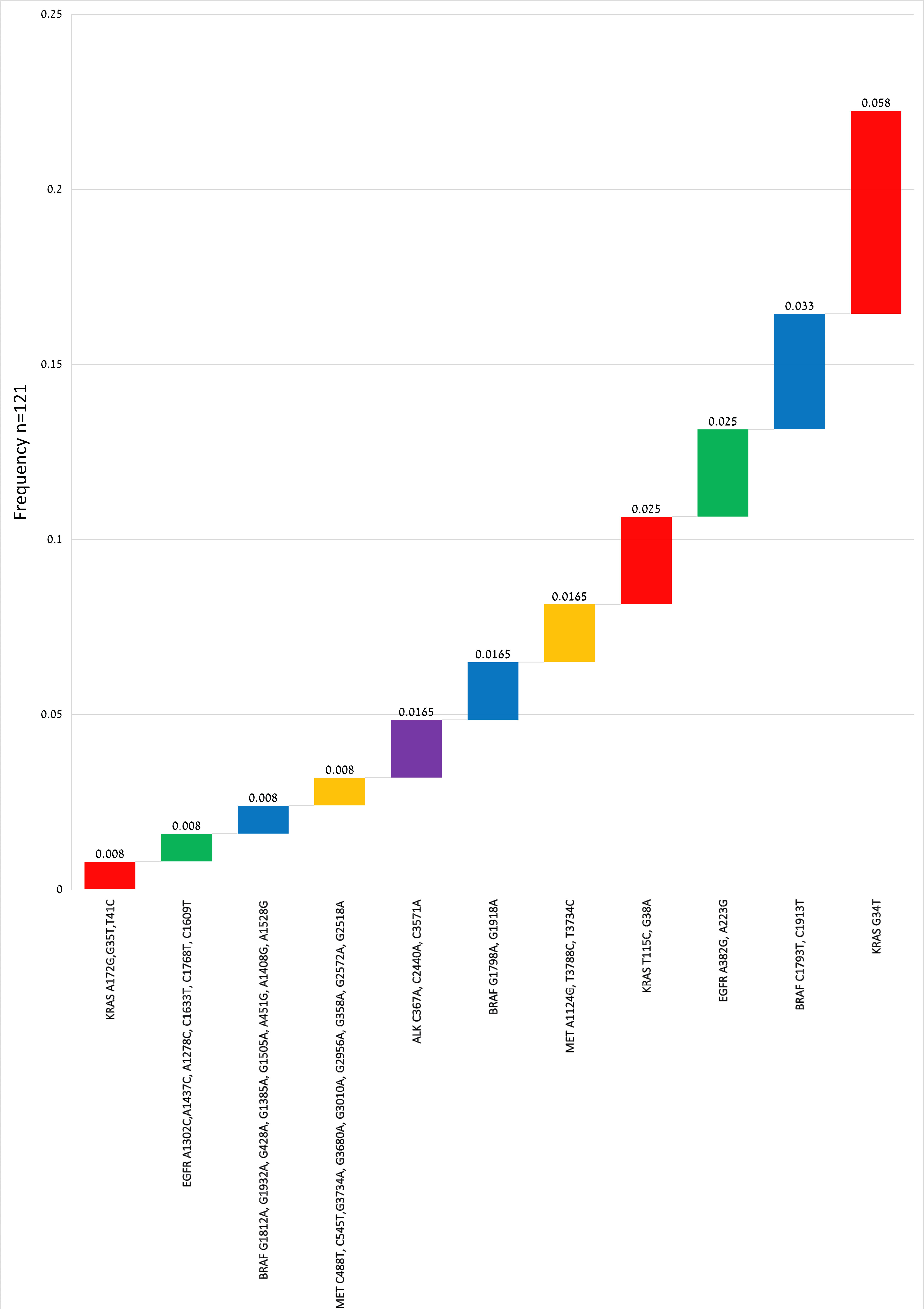
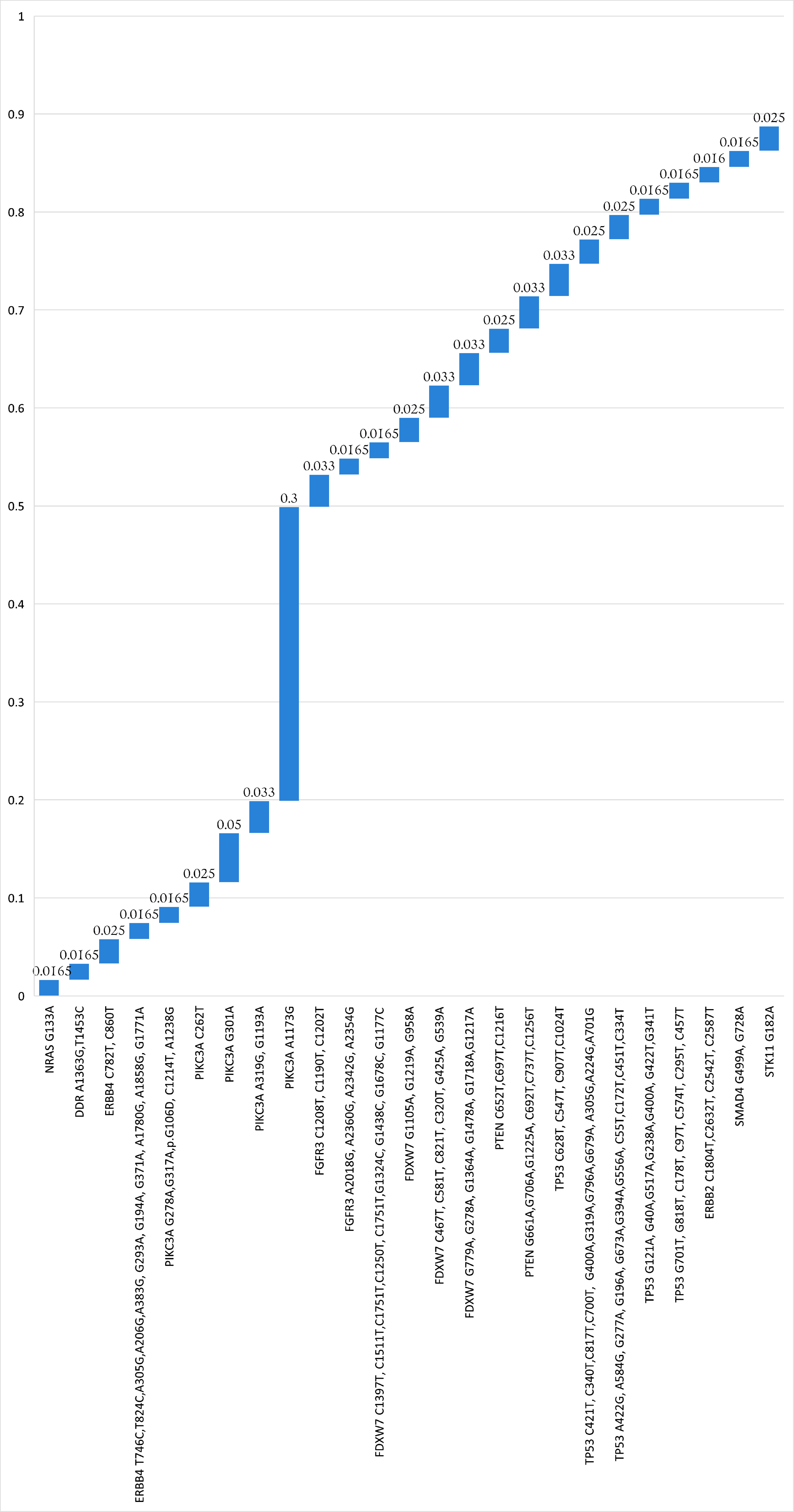
↓ Figure 1. The mutations for each gene including
synonymous variants (a), and excluding synonymous variants arranged in ascending order (b).
| World Journal of Oncology, ISSN 1920-4531 print, 1920-454X online, Open Access |
| Article copyright, the authors; Journal compilation copyright, World J Oncol and Elmer Press Inc |
| Journal website https://wjon.elmerpub.com |
Original Article
Volume 16, Number 2, April 2025, pages 161-172
Actionable Mutations and Survival Rates in Non-Small Cell Lung Cancer
Figures

Tables
| Clinical and pathological parameters | N (%) (n = 121) |
|---|---|
| Tumor histology | |
| Non-small cell lung cancer (NSCLC) | 107 (88) |
| Adenosquamous | 1 (0.8) |
| Large-cell neuroendocrine carcinoma | 1 (0.8) |
| Squamous cell carcinoma | 26 (21) |
| Adenocarcinoma | 79 (65) |
| Small cell lung cancer (SCLC) | 3 (2.4) |
| Undefined | 10 (8) |
| Gender | |
| Male | 101 (83) |
| Female | 20 (17) |
| Age | |
| ≤ 60 | 33 (27) |
| > 60 | 88 (73) |
| Clinical stage | |
| T1 | 31 (26) |
| T2 | 36 (30) |
| T3 | 38 (31) |
| T4 | 13 (11) |
| Metastasis | |
| Yes | 9 (7) |
| No | 89 (74) |
| Undefined | 23 (19) |
| Primary lung tumor | |
| Yes | 88 (73) |
| No | 10 (8) |
| Unknown | 23 (19) |
| Smoking | |
| Yes | 87 (72) |
| No | 21 (17) |
| Unknown | 13 (11) |
| Gene | Frameshift type | CDS change: AA change | Frequency (%) (n = 121) |
|---|---|---|---|
| CDS: coding sequence; AA: amino acid. | |||
| NRAS | Non-frameshift deletion | c.122_124del:p.R41del | 1 (0.008) |
| Non-frameshift insertion | c.26_27insGGT:p.V9_G10insV | 4 (0.033) | |
| DDR2 | Non-frameshift insertion | c.1449_1450insAAATAT:p.E483_P484insKY | 1 (0.008) |
| Non-frameshift deletion | c.1764_1769del:p.K591_D592del | 1 (0.008) | |
| Non-frameshift deletion | c.2343_2360del:p.F781_Q787delinsL | 2 (0.0165) | |
| ALK | Frameshift insertion | c.562_563insAGAAGCTGTC:p.L188Qfs*32 | 1 (0.008) |
| c.2635_2636insAGAAGCTGTC:p.L879Qfs*32 | |||
| c.3766_3767insAGAAGCTGTC:p.L1256Qfs*32 | |||
| ERBB4 | Frameshift insertion | c.1720dupG:p.A574Gfs*10 | 7 (0.058) |
| c.1798dupG:p.A600Gfs*10 | |||
| Frameshift deletion | c.720delT:p.F240Lfs*34 | 1 (0.008) | |
| c.798delT:p.F266Lfs*34 | |||
| Frameshift deletion | c.712delC:p.Q238Kfs*36 | 2 (0.0165) | |
| c.790delC:p.Q264Kfs*36 | |||
| Frameshift deletion | c.248_257del:p.K83Rfs*10 | 1 (0.008) | |
| c.326_335del:p.K109Rfs*10 | |||
| c.149_158del:p.K50Rfs*10 | |||
| PIKC3A | Stopgain | c.292delT:p.L99* | 2 (0.0165) |
| Stopgain | c.297delA:p.V101* | 1 (0.008) | |
| Frameshift deletion | c.1246delA:p.E417Rfs*11 | 2 (0.0165) | |
| Frameshift insertion | c.2686_2687insTG:p.F897Cfs*11 | 1 (0.008) | |
| FGFR3 | Frameshift insertion | c.2047dupC:p.A685Gfs*20 | 2 (0.0165) |
| c.2315dupC:p.G774Rfs*24 | |||
| c.2389dupC:p.A799Gfs*20 | |||
| Frameshift insertion | c.2068dupG:p.S692Lfs*13 | 2 (0.0165) | |
| c.2336dupG:p.L781Afs*17 | |||
| c.2410dupG:p.S806Lfs*13 | |||
| c.2392dupG:p.S800Lfs*13 | |||
| c.2404dupG:p.S804Lfs*13 | |||
| EGFR | Non-frameshift insertion | c.2161_2162insTGGCCAGCG:p.V724_D725insASV | 1 (0.008) |
| c.2296_2297insTGGCCAGCG:p.V769_D770insASV: | |||
| c.2137_2138insTGGCCAGCG:p.V716_D717insASV | |||
| FGFR1 | Non-frameshift insertion | c.121_122insGTG:p.D40_D41insG | 2 (0.0165) |
| c.121_122insGTG:p.D40_D41insG | |||
| c.388_389insGTG:p.D129_D130insG | |||
| Non-frameshift insertion | c.114_115insAAT:p.D38_D39insN | 2 (0.0165) | |
| c.381_382insAAT:p.D127_D128insN | |||
| c.114_115insAAT:p.D38_D39insN | |||
| NOTCH1 | Frameshift insertion | c.2331_2332insTG:p.L778Cfs*2 | 49 (0.40) |
| c.4620_4621insTG:p.L1541Cfs*2 | |||
| c.4734_4735insTG:p.L1579Cfs*2 | |||
| c.4614_4615insTG:p.L1539Cfs*2 | |||
| PTEN | Frameshift deletion | c.668_672del:p.F223Lfs*3 | 1 (0.008) |
| c.713_717del:p.F238Lfs*3 | |||
| c.1232_1236del:p.F411Lfs*3 | |||
| Frameshift deletion | c.750delA:p.K252Rfs*9 | 3 (0.025) | |
| c.795delA:p.K267Rfs*9 | |||
| c.1314delA:p.K440Rfs*9 | |||
| Frameshift insertion | c.910dupA:p.T304Nfs*6 | 1 (0.008) | |
| c.955dupA:p.T319Nfs*6 | |||
| c.1474dupA:p.T492Nfs*6 | |||
| Frameshift deletion | c.918delA:p.N308Mfs*21 | 3 (0.025) | |
| c.963delA:p.N323Mfs*40 | |||
| c.963delA:p.N323Mfs*21 | |||
| c.1482delA:p.N496Mfs*21 | |||
| FGFR2 | Frameshift insertion | c.765dupG:p.E256Gfs*66 | 2 (0.0165) |
| c.756dupG:p.E253Gfs*64 | |||
| c.759dupG:p.E254Gfs*66 | |||
| c.834dupG:p.E279Gfs*64 | |||
| c.1101dupG:p.E368Gfs*64 | |||
| c.834dupG:p.E279Gfs*66 | |||
| c.837dupG:p.E280Gfs*66 | |||
| c.1104dupG:p.E369Gfs*66 | |||
| c.417dupG:p.E140Gfs*66 | |||
| c.834dupG:p.E279Gfs*66 | |||
| c.1101dupG:p.E368Gfs*66 | |||
| c.1104dupG:p.E369Gfs*66 | |||
| AKT1 | Non-frameshift insertion | c.87_88insCTC:p.L29_K30insL | 1 (0.008) |
| TP53 | Frameshift insertion | c.398dupT:p.R135Tfs*5 | 1 (0.008) |
| c.317dupT:p.R108Tfs*5 | |||
| c.794dupT:p.R267Tfs*5 | |||
| c.677dupT:p.R228Tfs*5 | |||
| Frameshift deletion | c.87delC:p.I30Sfs*8 | 1 (0.008) | |
| c.6delC:p.I3Sfs*8 | |||
| c.87delC:p.I30Sfs*8 | |||
| c.483delC:p.I162Sfs*8 | |||
| c.204delC:p.I69Sfs*8 | |||
| c.366delC:p.I123Sfs*8 | |||
| SMAD4 | Non-frameshift insertion | c.326_327insAAAACATGA:p.H111_V112insEKH | 1 (0.008) |
| Frameshift insertion | c.397_398insACGA:p.V136Tfs*8 | 2 (0.0165) | |
| c.397_398insACGA:p.V136Tfs*6 | |||
| Frameshift insertion | c.936dupT:p.P313Sfs*46 | 2 (0.0165) | |
| c.936dupT:p.P313Sfs*9 | |||
| Clinical and pathological parameters | Carriers of actionable mutationsa (n = 35) | Non-carriers of actionable mutations (n = 86) | P valueb |
|---|---|---|---|
| aExcluding synonymous mutations. bBased on carriers of one common mutation or more. Chi-squared test or Fisher’s exact test was used (excluding missing data). The result is not significant (P > 0.05). | |||
| Tumor histology | 0.574 | ||
| Small cell lung cancer (SCLC) | 0 | 3 | |
| NSCLC (adenosquamous) | 0 | 1 | |
| NSCLC (large-cell neuroendocrine carcinoma) | 0 | 0 | |
| NSCLC (squamous cell carcinoma) | 5 | 18 | |
| NSCLC (adenocarcinoma) | 23 | 56 | |
| Unknown | 7 | 7 | |
| Gender | 0.752 | ||
| Male | 27 | 66 | |
| Female | 8 | 19 | |
| Age | 0.946 | ||
| ≤ 60 | 7 | 20 | |
| > 60 | 21 | 58 | |
| Unknown | 7 | 9 | |
| Tumor clinical stage | 0.438 | ||
| T1 | 7 | 20 | |
| T2 | 11 | 22 | |
| T3 | 11 | 22 | |
| T4 | 6 | 4 | |
| Metastasis | 0.797 | ||
| Yes | 3 | 9 | |
| No | 24 | 60 | |
| Unknown | 8 | 17 | |
| Primary tumor | 0.774 | ||
| Yes | 24 | 69 | |
| No | 3 | 7 | |
| Unknown | 8 | 10 | |
| Smoking | 0.484 | ||
| Yes | 21 | 51 | |
| No | 2 | 9 | |
| Unknown | 13 | 27 | |